You are browsing environment: FUNGIDB
CAZyme Information: ODM22814.1
You are here: Home > Sequence: ODM22814.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus cristatus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus cristatus | |||||||||||
| CAZyme ID | ODM22814.1 | |||||||||||
| CAZy Family | GT90 | |||||||||||
| CAZyme Description | UDP-N-acetylglucosamine transferase subunit alg13 | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 397977 | Glyco_tran_28_C | 6.19e-18 | 3 | 145 | 1 | 121 | Glycosyltransferase family 28 C-terminal domain. The glycosyltransferase family 28 includes monogalactosyldiacylglycerol synthase (EC 2.4.1.46) and UDP-N-acetylglucosamine transferase (EC 2.4.1.-). Structural analysis suggests the C-terminal domain contains the UDP-GlcNAc binding site. |
| 227350 | COG5017 | 2.71e-15 | 5 | 188 | 3 | 157 | UDP-N-acetylglucosamine transferase subunit ALG13 [Carbohydrate transport and metabolism]. |
| 223779 | MurG | 3.36e-09 | 83 | 168 | 246 | 334 | UDP-N-acetylglucosamine:LPS N-acetylglucosamine transferase [Cell wall/membrane/envelope biogenesis]. |
| 340818 | GT28_MurG | 4.58e-08 | 21 | 143 | 195 | 300 | undecaprenyldiphospho-muramoylpentapeptide beta-N-acetylglucosaminyltransferase. MurG (EC 2.4.1.227) is an N-acetylglucosaminyltransferase, the last enzyme involved in the intracellular phase of peptidoglycan biosynthesis. It transfers N-acetyl-D-glucosamine (GlcNAc) from UDP-GlcNAc to the C4 hydroxyl of a lipid-linked N-acetylmuramoyl pentapeptide (NAM). The resulting disaccharide is then transported across the cell membrane, where it is polymerized into NAG-NAM cell-wall repeat structure. MurG belongs to the GT-B structural superfamily of glycoslytransferases, which have characteristic N- and C-terminal domains, each containing a typical Rossmann fold. The two domains have high structural homology despite minimal sequence homology. The large cleft that separates the two domains includes the catalytic center and permits a high degree of flexibility. |
| 224732 | YjiC | 2.84e-05 | 2 | 163 | 238 | 363 | UDP:flavonoid glycosyltransferase YjiC, YdhE family [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 6.29e-136 | 1 | 194 | 1 | 194 | |
| 8.25e-86 | 1 | 194 | 10 | 204 | |
| 8.25e-86 | 1 | 194 | 10 | 204 | |
| 8.25e-86 | 1 | 194 | 10 | 204 | |
| 8.25e-86 | 1 | 194 | 10 | 204 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 9.48e-18 | 2 | 190 | 6 | 199 | NMR solution structure of ALG13 --- obtained with iterative CS-Rosetta from backbone NMR data. [Saccharomyces cerevisiae] |
|
| 1.45e-17 | 2 | 190 | 29 | 222 | NMR solution structure of ALG13: The sugar donor subunit of a yeast N-acetylglucosamine transferase. Northeast Structural Genomics Consortium target YG1 [Saccharomyces cerevisiae] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.19e-77 | 2 | 194 | 5 | 197 | UDP-N-acetylglucosamine transferase subunit alg13 OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=alg13 PE=3 SV=2 |
|
| 6.15e-21 | 5 | 158 | 4 | 136 | UDP-N-acetylglucosamine transferase subunit alg13 OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=alg13 PE=3 SV=1 |
|
| 1.90e-20 | 3 | 193 | 29 | 196 | UDP-N-acetylglucosamine transferase subunit ALG13 OS=Yarrowia lipolytica (strain CLIB 122 / E 150) OX=284591 GN=ALG13 PE=3 SV=1 |
|
| 5.71e-20 | 6 | 192 | 9 | 199 | UDP-N-acetylglucosamine transferase subunit ALG13 OS=Cryptococcus neoformans var. neoformans serotype D (strain JEC21 / ATCC MYA-565) OX=214684 GN=ALG13 PE=3 SV=1 |
|
| 5.71e-20 | 6 | 192 | 9 | 199 | UDP-N-acetylglucosamine transferase subunit ALG13 OS=Cryptococcus neoformans var. neoformans serotype D (strain B-3501A) OX=283643 GN=ALG13 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
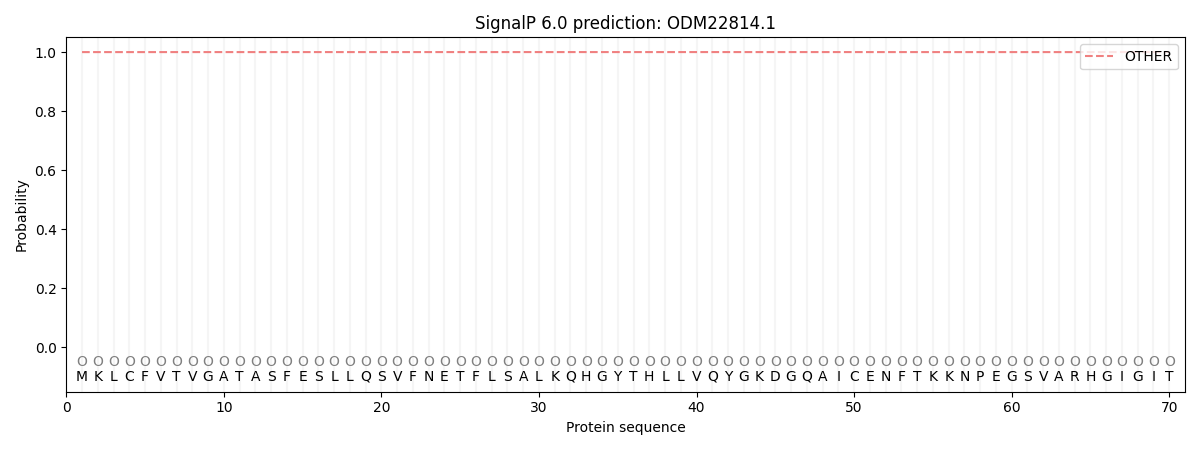
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000054 | 0.000003 |
