You are browsing environment: FUNGIDB
CAZyme Information: ODM21381.1
You are here: Home > Sequence: ODM21381.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
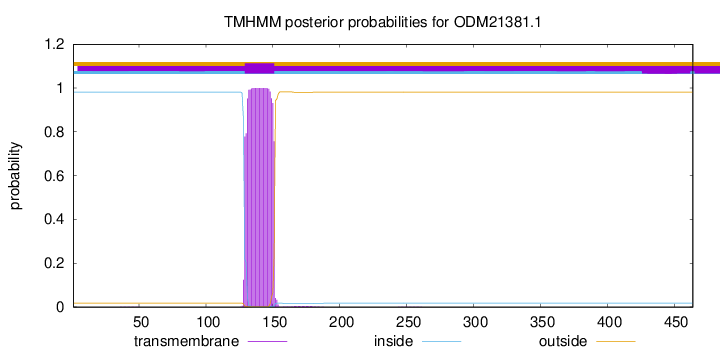
TMHMM annotations
Basic Information help
| Species | Aspergillus cristatus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus cristatus | |||||||||||
| CAZyme ID | ODM21381.1 | |||||||||||
| CAZy Family | GT1 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT34 | 163 | 413 | 4.7e-84 | 0.983739837398374 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 368535 | Glyco_transf_34 | 3.93e-84 | 163 | 412 | 3 | 238 | galactosyl transferase GMA12/MNN10 family. This family contains a number of glycosyltransferase enzymes that contain a DXD motif. This family includes a number of C. elegans homologs where the DXD is replaced by DXH. Some members of this family are included in glycosyltransferase family 34. |
| 215617 | PLN03182 | 5.69e-10 | 157 | 418 | 121 | 379 | xyloglucan 6-xylosyltransferase; Provisional |
| 215616 | PLN03181 | 6.02e-08 | 195 | 360 | 156 | 309 | glycosyltransferase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 464 | 1 | 464 | |
| 2.73e-177 | 1 | 456 | 1 | 463 | |
| 2.73e-177 | 1 | 446 | 1 | 456 | |
| 2.73e-177 | 1 | 456 | 1 | 463 | |
| 2.73e-177 | 1 | 446 | 1 | 456 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.45e-08 | 152 | 331 | 12 | 172 | Chain A, Xyloglucan 6-xylosyltransferase 1 [Arabidopsis thaliana],6BSU_B Chain B, Xyloglucan 6-xylosyltransferase 1 [Arabidopsis thaliana],6BSV_A Chain A, Xyloglucan 6-xylosyltransferase 1 [Arabidopsis thaliana],6BSV_B Chain B, Xyloglucan 6-xylosyltransferase 1 [Arabidopsis thaliana],6BSW_A Chain A, Xyloglucan 6-xylosyltransferase 1 [Arabidopsis thaliana],6BSW_B Chain B, Xyloglucan 6-xylosyltransferase 1 [Arabidopsis thaliana] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.39e-102 | 155 | 420 | 112 | 376 | Probable alpha-1,6-mannosyltransferase MNN10 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=MNN10 PE=1 SV=1 |
|
| 3.74e-93 | 155 | 421 | 63 | 332 | Uncharacterized alpha-1,2-galactosyltransferase C637.06 OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=SPAC637.06 PE=2 SV=2 |
|
| 7.20e-28 | 164 | 419 | 103 | 346 | Uncharacterized alpha-1,2-galactosyltransferase C1289.13c OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=SPBC1289.13c PE=3 SV=1 |
|
| 2.53e-27 | 164 | 420 | 103 | 347 | Alpha-1,2-galactosyltransferase OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=gma12 PE=1 SV=1 |
|
| 1.02e-25 | 195 | 412 | 114 | 320 | Alpha-1,2-galactosyltransferase gmh3 OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=gmh3 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER

| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000027 | 0.000016 |