You are browsing environment: FUNGIDB
CAZyme Information: OAG14986.1
You are here: Home > Sequence: OAG14986.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Alternaria alternata | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Dothideomycetes; ; Pleosporaceae; Alternaria; Alternaria alternata | |||||||||||
| CAZyme ID | OAG14986.1 | |||||||||||
| CAZy Family | AA7 | |||||||||||
| CAZyme Description | PMT-domain-containing protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 2.4.1.109:8 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT39 | 421 | 665 | 1.3e-72 | 0.9910313901345291 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 224839 | PMT1 | 0.0 | 413 | 1087 | 22 | 699 | Dolichyl-phosphate-mannose--protein O-mannosyl transferase [Posttranslational modification, protein turnover, chaperones]. |
| 406576 | PMT_4TMC | 7.74e-82 | 862 | 1079 | 1 | 197 | C-terminal four TMM region of protein-O-mannosyltransferase. PMT_4TMC is the C-terminal four membrane-pass region of protein-O-mannosyltransferases and similar enzymes. |
| 396786 | PMT | 1.94e-79 | 422 | 669 | 4 | 245 | Dolichyl-phosphate-mannose-protein mannosyltransferase. This is a family of Dolichyl-phosphate-mannose-protein mannosyltransferase proteins EC:2.4.1.109. These proteins are responsible for O-linked glycosylation of proteins, they catalyze the reaction:- Dolichyl phosphate D-mannose + protein <=> dolichyl phosphate + O-D-mannosyl-protein. Also in this family is Drosophila rotated abdomen protein which is a putative mannosyltransferase. This family appears to be distantly related to pfam02516 (A Bateman pers. obs.). This family also contains sequences from ArnTs (4-amino-4-deoxy-L-arabinose lipid A transferase). They catalyze the addition of 4-amino-4-deoxy-l-arabinose (l-Ara4N) to the lipid A moiety of the lipopolysaccharide. This is a critical modification enabling bacteria (e.g. Escherichia coli and Salmonella typhimurium) to resist killing by antimicrobial peptides such as polymyxins. Members such as undecaprenyl phosphate-alpha-4-amino-4-deoxy-L-arabinose arabinosyl transferase are predicted to have 12 trans-membrane regions. The N-terminal portion of these proteins is hypothesized to have a conserved glycosylation activity which is shared between distantly related oligosaccharyltransferases ArnT and PglB families. |
| 312076 | UAA | 3.46e-67 | 8 | 327 | 3 | 298 | UAA transporter family. This family includes transporters with a specificity for UDP-N-acetylglucosamine. |
| 129885 | nst | 7.19e-21 | 96 | 327 | 2 | 222 | UDP-galactose transporter. The 10-12 TMS Nucleotide Sugar Transporters (TC 2.A.7.10)Nucleotide-sugar transporters (NSTs) are found in the Golgi apparatus and the endoplasmic reticulum of eukaryotic cells. Members of the family have been sequenced from yeast, protozoans and animals. Animals such as C. elegans possess many of these transporters. Humans have at least two closely related isoforms of the UDP-galactose:UMP exchange transporter.NSTs generally appear to function by antiport mechanisms, exchanging a nucleotide-sugar for a nucleotide. Thus, CMP-sialic acid is exchanged for CMP; GDP-mannose is preferentially exchanged for GMP, and UDP-galactose and UDP-N-acetylglucosamine are exchanged for UMP (or possibly UDP). Other nucleotide sugars (e.g., GDP-fucose, UDP-xylose, UDP-glucose, UDP-N-acetylgalactosamine, etc.) may also be transported in exchange for various nucleotides, but their transporters have not been molecularly characterized. Each compound appears to be translocated by its own transport protein. Transport allows the compound, synthesized in the cytoplasm, to be exported to the lumen of the Golgi apparatus or the endoplasmic reticulum where it is used for the synthesis of glycoproteins and glycolipids. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 357 | 1087 | 1 | 807 | |
| 0.0 | 357 | 1087 | 1 | 771 | |
| 0.0 | 351 | 1087 | 48 | 817 | |
| 0.0 | 357 | 1087 | 1 | 771 | |
| 0.0 | 357 | 1087 | 1 | 779 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.47e-102 | 415 | 1082 | 50 | 728 | Structure of S. cerevisiae protein O-mannosyltransferase Pmt1-Pmt2 complex bound to the sugar donor and a peptide acceptor [Saccharomyces cerevisiae W303],6P2R_A Structure of S. cerevisiae protein O-mannosyltransferase Pmt1-Pmt2 complex bound to the sugar donor [Saccharomyces cerevisiae W303] |
|
| 1.16e-84 | 402 | 1067 | 52 | 737 | Structure of S. cerevisiae protein O-mannosyltransferase Pmt1-Pmt2 complex bound to the sugar donor and a peptide acceptor [Saccharomyces cerevisiae W303],6P2R_B Structure of S. cerevisiae protein O-mannosyltransferase Pmt1-Pmt2 complex bound to the sugar donor [Saccharomyces cerevisiae W303] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.20e-232 | 411 | 1087 | 44 | 755 | Dolichyl-phosphate-mannose--protein mannosyltransferase 4 OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=PMT4 PE=2 SV=2 |
|
| 3.67e-217 | 411 | 1087 | 49 | 762 | Dolichyl-phosphate-mannose--protein mannosyltransferase 4 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=PMT4 PE=1 SV=1 |
|
| 3.01e-211 | 410 | 1079 | 54 | 767 | Dolichyl-phosphate-mannose--protein mannosyltransferase 4 OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=ogm4 PE=3 SV=1 |
|
| 1.29e-106 | 413 | 1078 | 36 | 736 | Protein O-mannosyl-transferase 1 OS=Mus musculus OX=10090 GN=Pomt1 PE=1 SV=1 |
|
| 1.84e-105 | 413 | 1078 | 36 | 736 | Protein O-mannosyl-transferase 1 OS=Rattus norvegicus OX=10116 GN=Pomt1 PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER

| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000016 | 0.000000 |
TMHMM Annotations download full data without filtering help
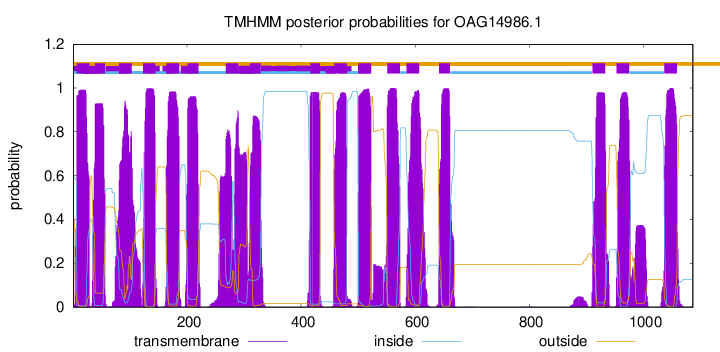
| Start | End |
|---|---|
| 7 | 29 |
| 39 | 56 |
| 81 | 103 |
| 123 | 144 |
| 164 | 186 |
| 201 | 220 |
| 268 | 290 |
| 310 | 329 |
| 416 | 433 |
| 458 | 480 |
| 501 | 523 |
| 551 | 573 |
| 585 | 607 |
| 642 | 661 |
| 911 | 933 |
| 953 | 975 |
| 1037 | 1059 |
