You are browsing environment: FUNGIDB
CAZyme Information: OAG14470.1
You are here: Home > Sequence: OAG14470.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Alternaria alternata | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Dothideomycetes; ; Pleosporaceae; Alternaria; Alternaria alternata | |||||||||||
| CAZyme ID | OAG14470.1 | |||||||||||
| CAZy Family | AA7 | |||||||||||
| CAZyme Description | glycoside hydrolase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH10 | 31 | 346 | 2.2e-100 | 0.9900990099009901 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 395262 | Glyco_hydro_10 | 3.13e-135 | 31 | 346 | 1 | 310 | Glycosyl hydrolase family 10. |
| 214750 | Glyco_10 | 3.70e-112 | 72 | 344 | 1 | 263 | Glycosyl hydrolase family 10. |
| 226217 | XynA | 8.15e-93 | 3 | 342 | 2 | 335 | Endo-1,4-beta-xylanase, GH35 family [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.96e-207 | 1 | 352 | 1 | 353 | |
| 4.44e-206 | 1 | 353 | 1 | 349 | |
| 3.07e-178 | 1 | 352 | 1 | 351 | |
| 4.76e-168 | 1 | 352 | 1 | 352 | |
| 1.18e-159 | 1 | 349 | 1 | 382 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7.43e-136 | 31 | 348 | 5 | 324 | A new crystal structure of a Fusarium oxysporum GH10 xylanase reveals the presence of an extended loop on top of the catalytic cleft [Fusarium oxysporum],3U7B_B A new crystal structure of a Fusarium oxysporum GH10 xylanase reveals the presence of an extended loop on top of the catalytic cleft [Fusarium oxysporum],3U7B_C A new crystal structure of a Fusarium oxysporum GH10 xylanase reveals the presence of an extended loop on top of the catalytic cleft [Fusarium oxysporum],3U7B_D A new crystal structure of a Fusarium oxysporum GH10 xylanase reveals the presence of an extended loop on top of the catalytic cleft [Fusarium oxysporum],3U7B_E A new crystal structure of a Fusarium oxysporum GH10 xylanase reveals the presence of an extended loop on top of the catalytic cleft [Fusarium oxysporum] |
|
| 1.54e-103 | 31 | 348 | 3 | 317 | Crystal structure of GH10 family xylanase XynAF1 from Aspergillus fumigatus Z5 [Aspergillus fumigatus Z5],6JDT_B Crystal structure of GH10 family xylanase XynAF1 from Aspergillus fumigatus Z5 [Aspergillus fumigatus Z5],6JDY_A Ligand complex structure of GH10 family xylanase XynAF1, soaking for 120 minutes [Aspergillus fumigatus Z5],6JDY_B Ligand complex structure of GH10 family xylanase XynAF1, soaking for 120 minutes [Aspergillus fumigatus Z5],6JDZ_A Ligand complex structure of GH10 family xylanase XynAF1, soaking for 20 minutes [Aspergillus fumigatus Z5],6JDZ_B Ligand complex structure of GH10 family xylanase XynAF1, soaking for 20 minutes [Aspergillus fumigatus Z5],6JE0_A Ligand complex structure of GH10 family xylanase XynAF1, soaking for 30 minutes [Aspergillus fumigatus Z5],6JE0_B Ligand complex structure of GH10 family xylanase XynAF1, soaking for 30 minutes [Aspergillus fumigatus Z5],6JE1_A Ligand complex structure of GH10 family xylanase XynAF1, soaking for 40 minutes [Aspergillus fumigatus Z5],6JE1_B Ligand complex structure of GH10 family xylanase XynAF1, soaking for 40 minutes [Aspergillus fumigatus Z5],6JE2_A Ligand complex structure of GH10 family xylanase XynAF1, soaking for 80 minutes [Aspergillus fumigatus Z5],6JE2_B Ligand complex structure of GH10 family xylanase XynAF1, soaking for 80 minutes [Aspergillus fumigatus Z5] |
|
| 4.82e-94 | 61 | 348 | 59 | 339 | GH10 endo-xylanase [Aspergillus aculeatus ATCC 16872],6Q8M_B GH10 endo-xylanase [Aspergillus aculeatus ATCC 16872],6Q8N_A GH10 endo-xylanase in complex with xylobiose epoxide inhibitor [Aspergillus aculeatus ATCC 16872],6Q8N_B GH10 endo-xylanase in complex with xylobiose epoxide inhibitor [Aspergillus aculeatus ATCC 16872] |
|
| 7.80e-81 | 52 | 341 | 27 | 305 | The mutant crystal structure of b-1,4-Xylanase (XynAS9_V43P/G44E) from Streptomyces sp. 9 [Streptomyces sp.],3WUG_A The mutant crystal structure of b-1,4-Xylanase (XynAS9_V43P/G44E) with xylobiose from Streptomyces sp. 9 [Streptomyces sp.] |
|
| 1.99e-78 | 52 | 341 | 27 | 305 | The wild type crystal structure of b-1,4-Xylanase (XynAS9) from Streptomyces sp. 9 [Streptomyces sp.],3WUE_A The wild type crystal structure of b-1,4-Xylanase (XynAS9) with xylobiose from Streptomyces sp. 9 [Streptomyces sp.] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5.39e-160 | 1 | 350 | 1 | 357 | Endo-1,4-beta-xylanase 2 OS=Aureobasidium pullulans OX=5580 GN=xynII PE=1 SV=1 |
|
| 3.82e-135 | 31 | 348 | 5 | 324 | Endo-1,4-beta-xylanase A OS=Fusarium oxysporum f. sp. lycopersici (strain 4287 / CBS 123668 / FGSC 9935 / NRRL 34936) OX=426428 GN=FOXG_17421 PE=1 SV=1 |
|
| 1.83e-132 | 26 | 350 | 16 | 342 | Endo-1,4-beta-xylanase 1 OS=Gibberella zeae (strain ATCC MYA-4620 / CBS 123657 / FGSC 9075 / NRRL 31084 / PH-1) OX=229533 GN=XYL1 PE=1 SV=1 |
|
| 1.25e-96 | 31 | 350 | 21 | 336 | Endo-1,4-beta-xylanase D OS=Talaromyces funiculosus OX=28572 GN=xynD PE=1 SV=1 |
|
| 7.11e-86 | 37 | 347 | 25 | 330 | Endo-1,4-beta-xylanase OS=Agaricus bisporus OX=5341 GN=xlnA PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
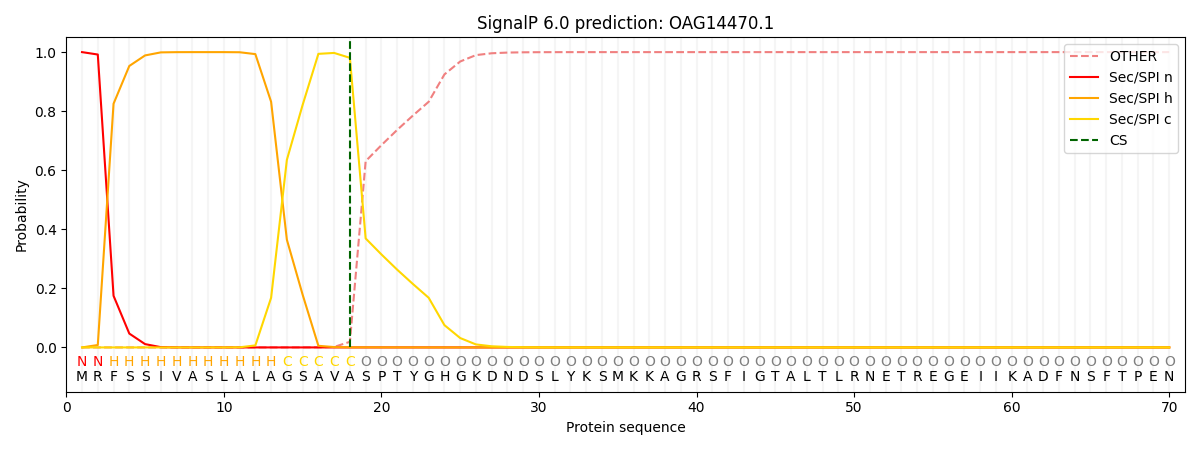
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000332 | 0.999618 | CS pos: 18-19. Pr: 0.9806 |
