You are browsing environment: FUNGIDB
CAZyme Information: OAG13697.1
You are here: Home > Sequence: OAG13697.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Alternaria alternata | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Dothideomycetes; ; Pleosporaceae; Alternaria; Alternaria alternata | |||||||||||
| CAZyme ID | OAG13697.1 | |||||||||||
| CAZy Family | AA3 | |||||||||||
| CAZyme Description | alpha/beta-hydrolase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE5 | 46 | 258 | 1.2e-36 | 0.9788359788359788 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 395860 | Cutinase | 7.74e-22 | 46 | 258 | 2 | 171 | Cutinase. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.09e-163 | 8 | 341 | 4 | 339 | |
| 1.91e-119 | 15 | 342 | 12 | 331 | |
| 9.47e-119 | 15 | 263 | 12 | 257 | |
| 1.11e-109 | 41 | 262 | 129 | 347 | |
| 8.50e-72 | 43 | 267 | 47 | 277 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.22e-08 | 46 | 256 | 2 | 202 | Acetylxylan Esterase From P. Purpurogenum Refined At 1.10 Angstroms [Talaromyces purpureogenus] |
|
| 1.22e-08 | 46 | 256 | 2 | 202 | ACETYLXYLAN ESTERASE AT 0.90 ANGSTROM RESOLUTION [Talaromyces purpureogenus] |
|
| 1.22e-08 | 46 | 256 | 2 | 202 | Iodinated Complex Of Acetyl Xylan Esterase At 1.80 Angstroms [Talaromyces purpureogenus] |
|
| 4.08e-08 | 46 | 256 | 2 | 202 | Chain A, ACETYL XYLAN ESTERASE [Trichoderma reesei],1QOZ_B Chain B, ACETYL XYLAN ESTERASE [Trichoderma reesei] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.17e-08 | 9 | 244 | 23 | 206 | Cutinase 4 OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=cut4 PE=2 SV=1 |
|
| 8.56e-08 | 46 | 256 | 29 | 229 | Acetylxylan esterase 2 OS=Talaromyces purpureogenus OX=1266744 GN=axe-2 PE=1 SV=1 |
|
| 2.47e-07 | 46 | 233 | 25 | 205 | Cutinase 11 OS=Verticillium dahliae OX=27337 GN=VD0003_g7577 PE=1 SV=1 |
|
| 2.57e-07 | 46 | 264 | 33 | 241 | Acetylxylan esterase OS=Hypocrea jecorina OX=51453 GN=axe1 PE=1 SV=1 |
|
| 9.12e-06 | 39 | 262 | 37 | 228 | Probable carboxylesterase Culp3 OS=Mycobacterium tuberculosis (strain CDC 1551 / Oshkosh) OX=83331 GN=cut3 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
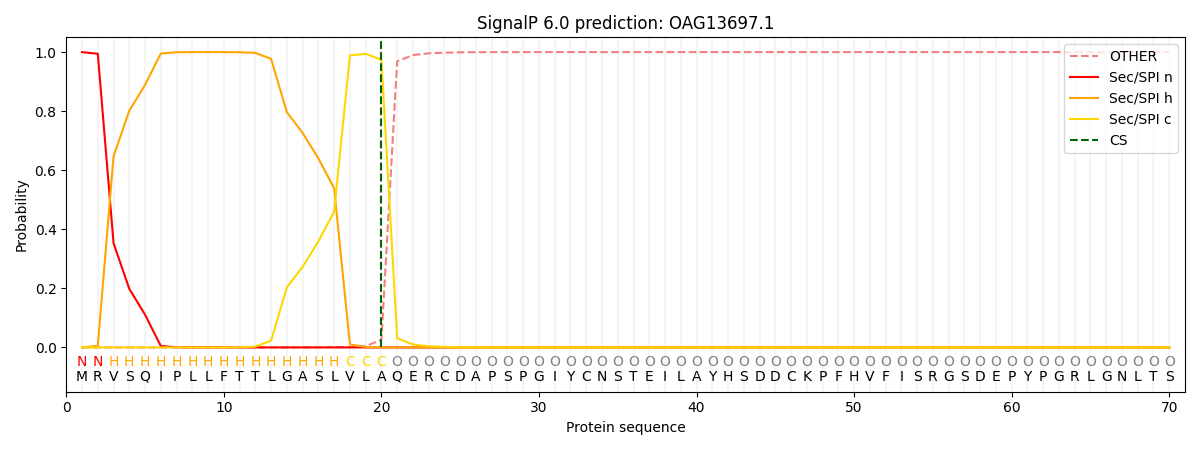
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000225 | 0.999743 | CS pos: 20-21. Pr: 0.9742 |
