You are browsing environment: FUNGIDB
CAZyme Information: NEUTE1DRAFT_131199-t26_1-p1
You are here: Home > Sequence: NEUTE1DRAFT_131199-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Neurospora tetrasperma | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Sordariaceae; Neurospora; Neurospora tetrasperma | |||||||||||
| CAZyme ID | NEUTE1DRAFT_131199-t26_1-p1 | |||||||||||
| CAZy Family | GH131 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 2.4.1.-:14 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT50 | 194 | 474 | 1.5e-98 | 0.9961832061068703 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 178434 | PLN02841 | 1.06e-77 | 46 | 474 | 28 | 415 | GPI mannosyltransferase |
| 252941 | Mannosyl_trans | 1.22e-73 | 194 | 474 | 1 | 259 | Mannosyltransferase (PIG-M). PIG-M has a DXD motif. The DXD motif is found in many glycosyltransferases that utilize nucleotide sugars. It is thought that the motif is involved in the binding of a manganese ion that is required for association of the enzymes with nucleotide sugar substrates. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.77e-208 | 9 | 476 | 10 | 441 | |
| 1.18e-206 | 9 | 476 | 10 | 441 | |
| 1.18e-206 | 9 | 476 | 10 | 441 | |
| 1.18e-206 | 9 | 476 | 10 | 441 | |
| 2.81e-187 | 7 | 476 | 2 | 426 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 0.0 | 1 | 479 | 1 | 487 | GPI mannosyltransferase 1 OS=Neurospora crassa (strain ATCC 24698 / 74-OR23-1A / CBS 708.71 / DSM 1257 / FGSC 987) OX=367110 GN=gim-1 PE=3 SV=1 |
|
| 1.01e-173 | 23 | 479 | 7 | 410 | GPI mannosyltransferase 1 OS=Gibberella zeae (strain ATCC MYA-4620 / CBS 123657 / FGSC 9075 / NRRL 31084 / PH-1) OX=229533 GN=GPI14 PE=3 SV=1 |
|
| 2.11e-171 | 19 | 476 | 1 | 409 | GPI mannosyltransferase 1 OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=gpi14 PE=3 SV=2 |
|
| 5.23e-169 | 19 | 478 | 1 | 429 | GPI mannosyltransferase 1 OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=gpi14 PE=3 SV=1 |
|
| 5.26e-167 | 19 | 478 | 1 | 422 | GPI mannosyltransferase 1 OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=gpi14 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER

| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000011 | 0.000003 |
TMHMM Annotations download full data without filtering help
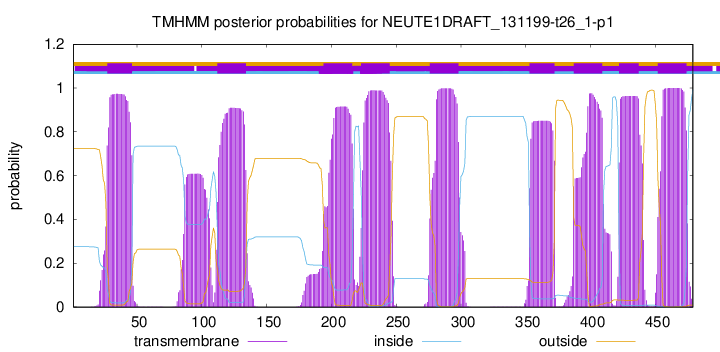
| Start | End |
|---|---|
| 27 | 46 |
| 112 | 134 |
| 194 | 216 |
| 223 | 245 |
| 276 | 298 |
| 353 | 372 |
| 387 | 409 |
| 422 | 437 |
| 452 | 474 |
