You are browsing environment: FUNGIDB
CAZyme Information: NEUTE1DRAFT_128956-t26_1-p1
You are here: Home > Sequence: NEUTE1DRAFT_128956-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Neurospora tetrasperma | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Sordariaceae; Neurospora; Neurospora tetrasperma | |||||||||||
| CAZyme ID | NEUTE1DRAFT_128956-t26_1-p1 | |||||||||||
| CAZy Family | GH114 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT59 | 216 | 532 | 6.4e-95 | 0.7054455445544554 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 398537 | DIE2_ALG10 | 1.36e-112 | 218 | 804 | 2 | 383 | DIE2/ALG10 family. The ALG10 protein from Saccharomyces cerevisiae encodes the alpha-1,2 glucosyltransferase of the endoplasmic reticulum. This protein has been characterized in rat as potassium channel regulator 1. |
| 401668 | QCR10 | 4.88e-19 | 28 | 92 | 1 | 61 | Ubiquinol-cytochrome-c reductase complex subunit (QCR10). The QCR10 family of proteins are a component of the ubiquinol-cytochrome c reductase complex (also known as complex III or cytochrome b-c1 complex). This complex is located on the inner mitochondrial membrane and it couples electron transfer from ubiquinol to cytochrome. This subunit (QCR10) is required for stable association of the iron-sulfur protein with the complex. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 2.71e-201 | 150 | 909 | 1 | 663 | |
| 3.53e-193 | 194 | 909 | 4 | 551 | |
| 2.80e-188 | 186 | 909 | 26 | 713 | |
| 8.41e-184 | 208 | 909 | 48 | 712 | |
| 1.26e-183 | 208 | 909 | 48 | 714 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 0.0 | 149 | 909 | 1 | 771 | Dol-P-Glc:Glc(2)Man(9)GlcNAc(2)-PP-Dol alpha-1,2-glucosyltransferase OS=Neurospora crassa (strain ATCC 24698 / 74-OR23-1A / CBS 708.71 / DSM 1257 / FGSC 987) OX=367110 GN=alg-10 PE=3 SV=3 |
|
| 1.85e-172 | 201 | 909 | 59 | 722 | Dol-P-Glc:Glc(2)Man(9)GlcNAc(2)-PP-Dol alpha-1,2-glucosyltransferase OS=Gibberella zeae (strain ATCC MYA-4620 / CBS 123657 / FGSC 9075 / NRRL 31084 / PH-1) OX=229533 GN=ALG10 PE=3 SV=1 |
|
| 1.82e-149 | 178 | 909 | 29 | 660 | Dol-P-Glc:Glc(2)Man(9)GlcNAc(2)-PP-Dol alpha-1,2-glucosyltransferase OS=Magnaporthe oryzae (strain 70-15 / ATCC MYA-4617 / FGSC 8958) OX=242507 GN=ALG10 PE=3 SV=1 |
|
| 7.58e-80 | 209 | 909 | 28 | 614 | Dol-P-Glc:Glc(2)Man(9)GlcNAc(2)-PP-Dol alpha-1,2-glucosyltransferase OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=alg10 PE=3 SV=1 |
|
| 2.88e-74 | 209 | 909 | 27 | 608 | Dol-P-Glc:Glc(2)Man(9)GlcNAc(2)-PP-Dol alpha-1,2-glucosyltransferase OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=alg10 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
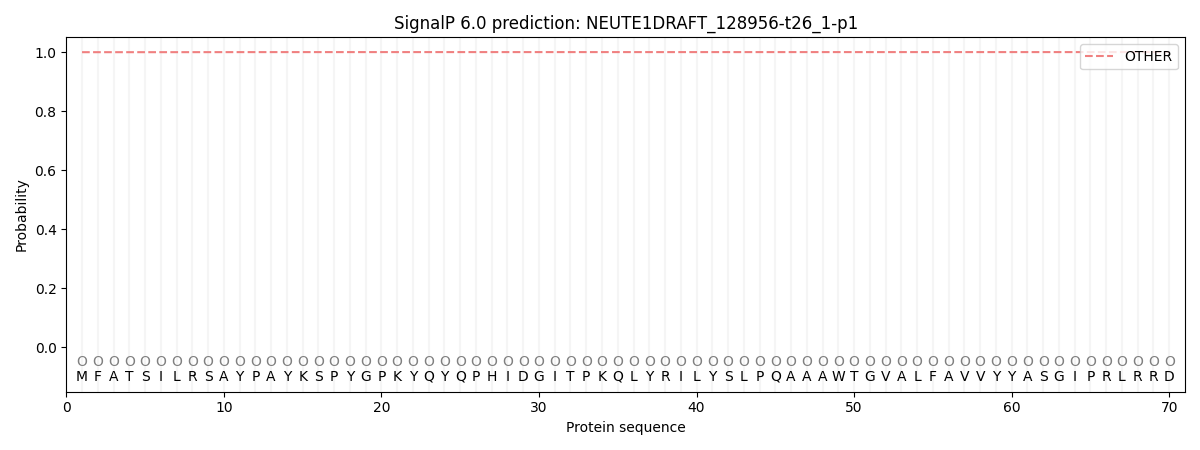
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.999865 | 0.000156 |
TMHMM Annotations download full data without filtering help
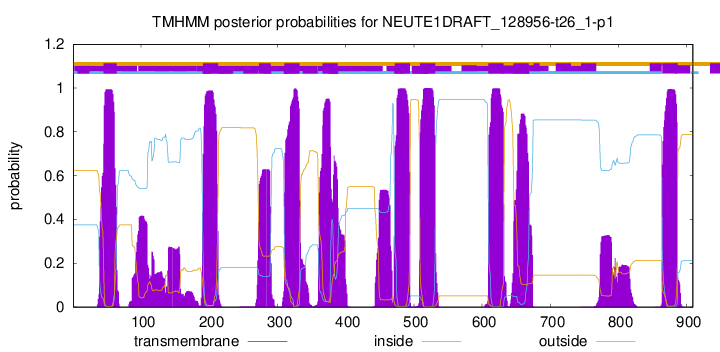
| Start | End |
|---|---|
| 45 | 64 |
| 191 | 213 |
| 273 | 290 |
| 310 | 332 |
| 366 | 388 |
| 472 | 494 |
| 509 | 531 |
| 609 | 631 |
| 646 | 668 |
| 864 | 886 |
