You are browsing environment: FUNGIDB
CAZyme Information: NEUTE1DRAFT_125708-t26_1-p1
You are here: Home > Sequence: NEUTE1DRAFT_125708-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Neurospora tetrasperma | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Sordariaceae; Neurospora; Neurospora tetrasperma | |||||||||||
| CAZyme ID | NEUTE1DRAFT_125708-t26_1-p1 | |||||||||||
| CAZy Family | CE9 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.28:1 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH37 | 53 | 628 | 1.7e-161 | 0.9918533604887984 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 395961 | Trehalase | 6.42e-137 | 53 | 631 | 3 | 509 | Trehalase. Trehalase (EC:3.2.1.28) is known to recycle trehalose to glucose. Trehalose is a physiological hallmark of heat-shock response in yeast and protects of proteins and membranes against a variety of stresses. This family is found in conjunction with pfam07492 in fungi. |
| 215307 | PLN02567 | 1.95e-125 | 51 | 631 | 25 | 545 | alpha,alpha-trehalase |
| 224541 | TreA | 3.99e-92 | 35 | 636 | 53 | 558 | Neutral trehalase [Carbohydrate transport and metabolism]. |
| 183934 | treF | 3.17e-80 | 40 | 622 | 59 | 538 | alpha,alpha-trehalase TreF. |
| 183936 | treA | 4.51e-78 | 1 | 631 | 11 | 537 | alpha,alpha-trehalase TreA. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 692 | 1 | 692 | |
| 0.0 | 14 | 675 | 13 | 678 | |
| 0.0 | 14 | 675 | 12 | 677 | |
| 0.0 | 13 | 673 | 19 | 700 | |
| 0.0 | 13 | 673 | 19 | 700 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.54e-68 | 55 | 631 | 40 | 555 | Chain A, Trehalase [Arabidopsis thaliana],7E9U_B Chain B, Trehalase [Arabidopsis thaliana] |
|
| 2.13e-67 | 55 | 631 | 40 | 555 | Chain A, Trehalase [Arabidopsis thaliana],7E9X_B Chain B, Trehalase [Arabidopsis thaliana],7E9X_C Chain C, Trehalase [Arabidopsis thaliana],7E9X_D Chain D, Trehalase [Arabidopsis thaliana],7EAW_A Chain A, Trehalase [Arabidopsis thaliana],7EAW_B Chain B, Trehalase [Arabidopsis thaliana] |
|
| 3.35e-66 | 35 | 638 | 46 | 547 | Structure of periplasmic trehalase from Diamondback moth gut bacteria complexed with validoxylamine [Enterobacter cloacae],5Z6H_A Structure of periplasmic trehalase from Diamondback moth gut bacteria in the apo form [Enterobacter cloacae],5Z6H_B Structure of periplasmic trehalase from Diamondback moth gut bacteria in the apo form [Enterobacter cloacae] |
|
| 1.79e-65 | 36 | 628 | 10 | 500 | Family 37 trehalase from Escherichia coli in complex with 1- thiatrehazolin [Escherichia coli K-12],2JJB_A Family 37 trehalase from Escherichia coli in complex with casuarine-6- O-alpha-glucopyranose [Escherichia coli K-12],2JJB_B Family 37 trehalase from Escherichia coli in complex with casuarine-6- O-alpha-glucopyranose [Escherichia coli K-12],2JJB_C Family 37 trehalase from Escherichia coli in complex with casuarine-6- O-alpha-glucopyranose [Escherichia coli K-12],2JJB_D Family 37 trehalase from Escherichia coli in complex with casuarine-6- O-alpha-glucopyranose [Escherichia coli K-12],2WYN_A Structure of family 37 trehalase from Escherichia coli in complex with a casuarine-6-O-a-D-glucoside analogue [Escherichia coli K-12],2WYN_B Structure of family 37 trehalase from Escherichia coli in complex with a casuarine-6-O-a-D-glucoside analogue [Escherichia coli K-12],2WYN_C Structure of family 37 trehalase from Escherichia coli in complex with a casuarine-6-O-a-D-glucoside analogue [Escherichia coli K-12],2WYN_D Structure of family 37 trehalase from Escherichia coli in complex with a casuarine-6-O-a-D-glucoside analogue [Escherichia coli K-12] |
|
| 6.39e-62 | 36 | 628 | 10 | 500 | Family 37 trehalase from Escherichia coli in complex with validoxylamine [Escherichia coli K-12] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.11e-130 | 20 | 639 | 36 | 594 | Trehalase OS=Dictyostelium discoideum OX=44689 GN=treh PE=3 SV=1 |
|
| 1.05e-118 | 32 | 634 | 24 | 551 | Trehalase OS=Mus musculus OX=10090 GN=Treh PE=1 SV=1 |
|
| 4.09e-115 | 32 | 634 | 27 | 553 | Trehalase OS=Oryctolagus cuniculus OX=9986 GN=TREH PE=1 SV=1 |
|
| 6.83e-113 | 24 | 638 | 32 | 576 | Trehalase OS=Apis mellifera OX=7460 PE=1 SV=1 |
|
| 1.12e-112 | 32 | 634 | 27 | 554 | Trehalase OS=Homo sapiens OX=9606 GN=TREH PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
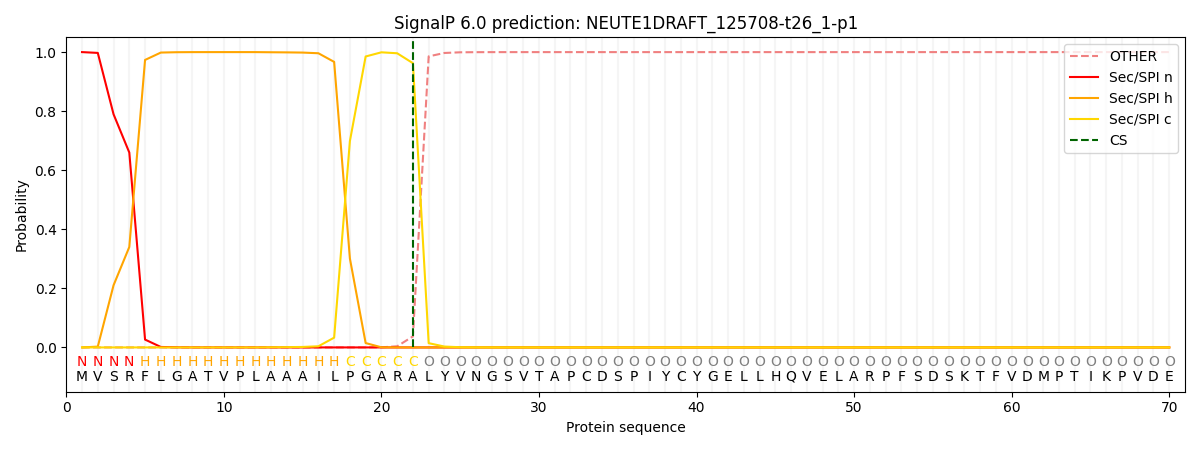
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000272 | 0.999705 | CS pos: 22-23. Pr: 0.9625 |
