You are browsing environment: FUNGIDB
CAZyme Information: NEUDI_63398T0-p1
You are here: Home > Sequence: NEUDI_63398T0-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
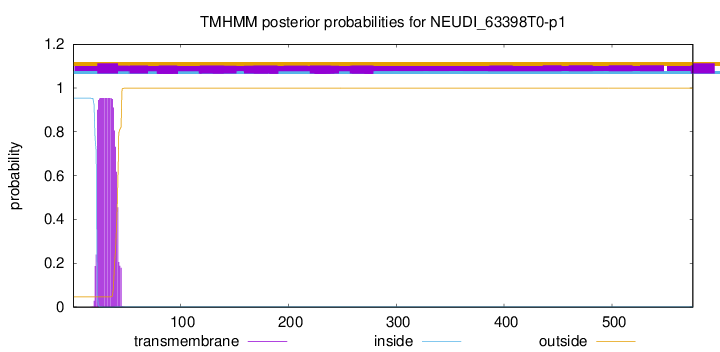
TMHMM annotations
Basic Information help
| Species | Neurospora discreta | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Sordariaceae; Neurospora; Neurospora discreta | |||||||||||
| CAZyme ID | NEUDI_63398T0-p1 | |||||||||||
| CAZy Family | GH7 | |||||||||||
| CAZyme Description | Mannosyl-oligosaccharide alpha-1,2-mannosidase and related glycosyl hydrolases | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.113:7 | 3.2.1.113:7 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH47 | 135 | 359 | 5.7e-71 | 0.49327354260089684 |
| GH47 | 353 | 563 | 3.6e-43 | 0.3968609865470852 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 396217 | Glyco_hydro_47 | 7.62e-149 | 134 | 558 | 1 | 416 | Glycosyl hydrolase family 47. Members of this family are alpha-mannosidases that catalyze the hydrolysis of the terminal 1,2-linked alpha-D-mannose residues in the oligo-mannose oligosaccharide Man(9)(GlcNAc)(2). |
| 240427 | PTZ00470 | 1.75e-87 | 125 | 544 | 70 | 469 | glycoside hydrolase family 47 protein; Provisional |
| 224250 | YyaL | 2.52e-04 | 121 | 373 | 393 | 626 | Uncharacterized conserved protein YyaL, SSP411 family, contains thoiredoxin and six-hairpin glycosidase-like domains [General function prediction only]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 4.83e-289 | 1 | 558 | 1 | 553 | |
| 6.03e-212 | 16 | 558 | 7 | 552 | |
| 3.83e-207 | 16 | 558 | 5 | 549 | |
| 1.09e-204 | 16 | 558 | 5 | 551 | |
| 2.98e-204 | 17 | 558 | 3 | 550 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5.45e-45 | 130 | 547 | 8 | 421 | Penicillium citrinum alpha-1,2-mannosidase complex with glycerol [Penicillium citrinum],2RI8_B Penicillium citrinum alpha-1,2-mannosidase complex with glycerol [Penicillium citrinum],2RI9_A Penicillium citrinum alpha-1,2-mannosidase in complex with a substrate analog [Penicillium citrinum],2RI9_B Penicillium citrinum alpha-1,2-mannosidase in complex with a substrate analog [Penicillium citrinum] |
|
| 1.02e-44 | 130 | 547 | 43 | 456 | Structure of P. citrinum alpha 1,2-mannosidase reveals the basis for differences in specificity of the ER and Golgi Class I enzymes [Penicillium citrinum],1KKT_B Structure of P. citrinum alpha 1,2-mannosidase reveals the basis for differences in specificity of the ER and Golgi Class I enzymes [Penicillium citrinum],1KRE_A Structure Of P. Citrinum Alpha 1,2-Mannosidase Reveals The Basis For Differences In Specificity Of The Er And Golgi Class I Enzymes [Penicillium citrinum],1KRE_B Structure Of P. Citrinum Alpha 1,2-Mannosidase Reveals The Basis For Differences In Specificity Of The Er And Golgi Class I Enzymes [Penicillium citrinum],1KRF_A Structure Of P. Citrinum Alpha 1,2-mannosidase Reveals The Basis For Differences In Specificity Of The Er And Golgi Class I Enzymes [Penicillium citrinum],1KRF_B Structure Of P. Citrinum Alpha 1,2-mannosidase Reveals The Basis For Differences In Specificity Of The Er And Golgi Class I Enzymes [Penicillium citrinum] |
|
| 2.43e-44 | 127 | 554 | 5 | 410 | Crystal structure of the class I human endoplasmic reticulum 1,2-alpha-mannosidase and Man9GlcNAc2-PA complex [Homo sapiens] |
|
| 2.58e-44 | 127 | 554 | 5 | 410 | Crystal structure of the class I human endoplasmic reticulum 1,2-alpha-mannosidase T688A mutant and Thio-disaccharide substrate analog complex [Homo sapiens],5KK7_B Crystal structure of the class I human endoplasmic reticulum 1,2-alpha-mannosidase T688A mutant and Thio-disaccharide substrate analog complex [Homo sapiens] |
|
| 2.73e-44 | 127 | 554 | 10 | 415 | Crystal Structure Of Human Class I Alpha1,2-Mannosidase [Homo sapiens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.51e-48 | 116 | 547 | 28 | 455 | Probable mannosyl-oligosaccharide alpha-1,2-mannosidase 1B OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=mns1B PE=3 SV=1 |
|
| 1.03e-46 | 115 | 544 | 89 | 489 | Mannosyl-oligosaccharide 1,2-alpha-mannosidase MNS1 OS=Arabidopsis thaliana OX=3702 GN=MNS1 PE=1 SV=1 |
|
| 4.18e-46 | 110 | 548 | 26 | 456 | Probable mannosyl-oligosaccharide alpha-1,2-mannosidase ARB_00035 OS=Arthroderma benhamiae (strain ATCC MYA-4681 / CBS 112371) OX=663331 GN=ARB_00035 PE=1 SV=1 |
|
| 4.25e-46 | 125 | 547 | 38 | 456 | Mannosyl-oligosaccharide alpha-1,2-mannosidase 1B OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=mns1B PE=1 SV=1 |
|
| 4.25e-46 | 125 | 547 | 38 | 456 | Probable mannosyl-oligosaccharide alpha-1,2-mannosidase 1B OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=mns1B PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER

| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.999946 | 0.000054 |