You are browsing environment: FUNGIDB
CAZyme Information: NEUDI_104045T0-p1
You are here: Home > Sequence: NEUDI_104045T0-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Neurospora discreta | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Sordariaceae; Neurospora; Neurospora discreta | |||||||||||
| CAZyme ID | NEUDI_104045T0-p1 | |||||||||||
| CAZy Family | AA2|AA2 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT69 | 151 | 395 | 8.1e-75 | 0.9916317991631799 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 403053 | CAP59_mtransfer | 5.23e-94 | 151 | 390 | 1 | 225 | Cryptococcal mannosyltransferase 1. The capsule of pathogenic fungi is a complex polysaccharide whose formation is determined by a number of enzymes including, most importantly, alpha-1,3-mannosyltransferase 1, EC:2.4.1.-. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.39e-242 | 1 | 472 | 1 | 457 | |
| 1.84e-184 | 94 | 472 | 61 | 442 | |
| 9.22e-178 | 113 | 472 | 1 | 355 | |
| 5.30e-177 | 113 | 472 | 1 | 355 | |
| 5.30e-177 | 113 | 472 | 1 | 355 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5.76e-58 | 165 | 455 | 142 | 443 | Alpha-1,3-mannosyltransferase CMT1 OS=Cryptococcus neoformans var. neoformans serotype D (strain JEC21 / ATCC MYA-565) OX=214684 GN=CMT1 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
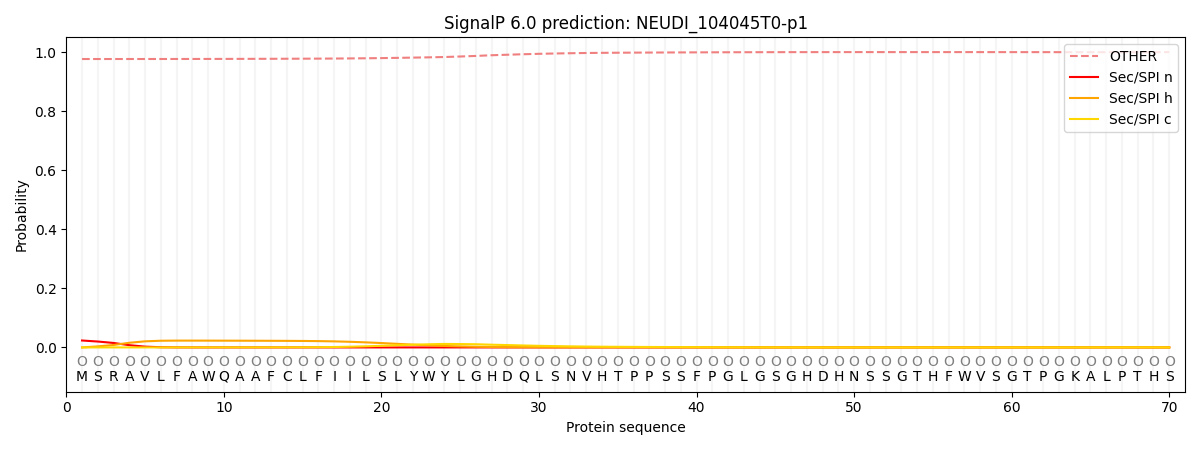
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.977814 | 0.022207 |