You are browsing environment: FUNGIDB
CAZyme Information: NCU12033-t26_1-p1
You are here: Home > Sequence: NCU12033-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
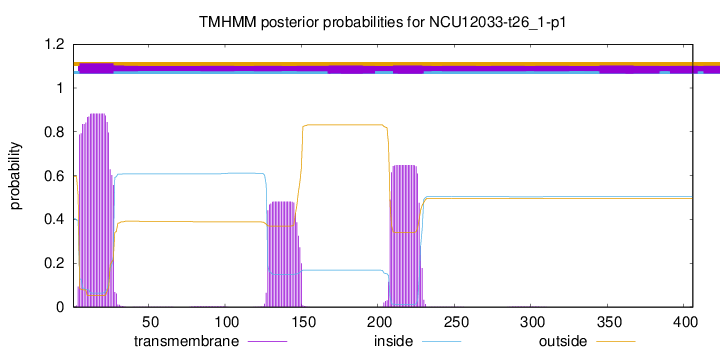
TMHMM annotations
Basic Information help
| Species | Neurospora crassa | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Sordariaceae; Neurospora; Neurospora crassa | |||||||||||
| CAZyme ID | NCU12033-t26_1-p1 | |||||||||||
| CAZy Family | GT69 | |||||||||||
| CAZyme Description | class III chitinase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.96:4 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH18 | 62 | 293 | 1.8e-17 | 0.6554054054054054 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 119363 | GH18_CTS3_chitinase | 6.68e-159 | 62 | 331 | 1 | 256 | GH18 domain of CTS3 (chitinase 3), an uncharacterized protein from the human fungal pathogen Coccidioides posadasii. CTS3 has a chitinase-like glycosyl hydrolase family 18 (GH18) domain; and has homologs in bacteria as well as fungi. |
| 119349 | GH18_chitinase-like | 1.79e-16 | 91 | 253 | 24 | 176 | The GH18 (glycosyl hydrolase, family 18) type II chitinases hydrolyze chitin, an abundant polymer of beta-1,4-linked N-acetylglucosamine (GlcNAc) which is a major component of the cell wall of fungi and the exoskeleton of arthropods. Chitinases have been identified in viruses, bacteria, fungi, protozoan parasites, insects, and plants. The structure of the GH18 domain is an eight-stranded beta/alpha barrel with a pronounced active-site cleft at the C-terminal end of the beta-barrel. The GH18 family includes chitotriosidase, chitobiase, hevamine, zymocin-alpha, narbonin, SI-CLP (stabilin-1 interacting chitinase-like protein), IDGF (imaginal disc growth factor), CFLE (cortical fragment-lytic enzyme) spore hydrolase, the type III and type V plant chitinases, the endo-beta-N-acetylglucosaminidases, and the chitolectins. The GH85 (glycosyl hydrolase, family 85) ENGases (endo-beta-N-acetylglucosaminidases) are closely related to the GH18 chitinases and are included in this alignment model. |
| 395573 | Glyco_hydro_18 | 2.26e-10 | 62 | 289 | 1 | 219 | Glycosyl hydrolases family 18. |
| 119350 | GH18_chitinase_D-like | 6.88e-08 | 132 | 326 | 72 | 293 | GH18 domain of Chitinase D (ChiD). ChiD, a chitinase found in Bacillus circulans, hydrolyzes the 1,4-beta-linkages of N-acetylglucosamine in chitin and chitodextrins. The domain architecture of ChiD includes a catalytic glycosyl hydrolase family 18 (GH18) domain, a chitin-binding domain, and a fibronectin type III domain. The chitin-binding and fibronectin type III domains are located either N-terminal or C-terminal to the catalytic domain. This family includes exochitinase Chi36 from Bacillus cereus. |
| 119353 | GH18_CFLE_spore_hydrolase | 8.35e-06 | 91 | 234 | 26 | 165 | Cortical fragment-lytic enzyme (CFLE) is a peptidoglycan hydrolase involved in bacterial endospore germination. CFLE is expressed as an inactive preprotein (called SleB) in the forespore compartment of sporulating cells. SleB translocates across the forespore inner membrane and is deposited as a mature enzyme in the cortex layer of the spore. As part of a sensory mechanism capable of initiating germination, CFLE degrades a spore-specific peptidoglycan constituent called muramic-acid delta-lactam that comprises the outer cortex. CFLE has a C-terminal glycosyl hydrolase family 18 (GH18) catalytic domain as well as two N-terminal LysM peptidoglycan-binding domains. In addition to SleB, this family includes YaaH, YdhD, and YvbX from Bacillus subtilis. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 3.04e-125 | 59 | 405 | 28 | 355 | |
| 4.28e-123 | 58 | 406 | 29 | 358 | |
| 1.83e-120 | 58 | 406 | 32 | 362 | |
| 2.59e-120 | 58 | 406 | 32 | 362 | |
| 4.07e-120 | 10 | 406 | 46 | 418 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.58e-110 | 59 | 353 | 1 | 282 | Chain X, ENDO-N-ACETYL-BETA-D-GLUCOSAMINIDASE [Trichoderma reesei] |
|
| 1.14e-70 | 57 | 347 | 9 | 285 | Structure of chitinase, ChiC, from Aspergillus fumigatus. [Aspergillus fumigatus Af293],2Y8V_B Structure of chitinase, ChiC, from Aspergillus fumigatus. [Aspergillus fumigatus Af293],2Y8V_C Structure of chitinase, ChiC, from Aspergillus fumigatus. [Aspergillus fumigatus Af293],2Y8V_D Structure of chitinase, ChiC, from Aspergillus fumigatus. [Aspergillus fumigatus Af293] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as SP
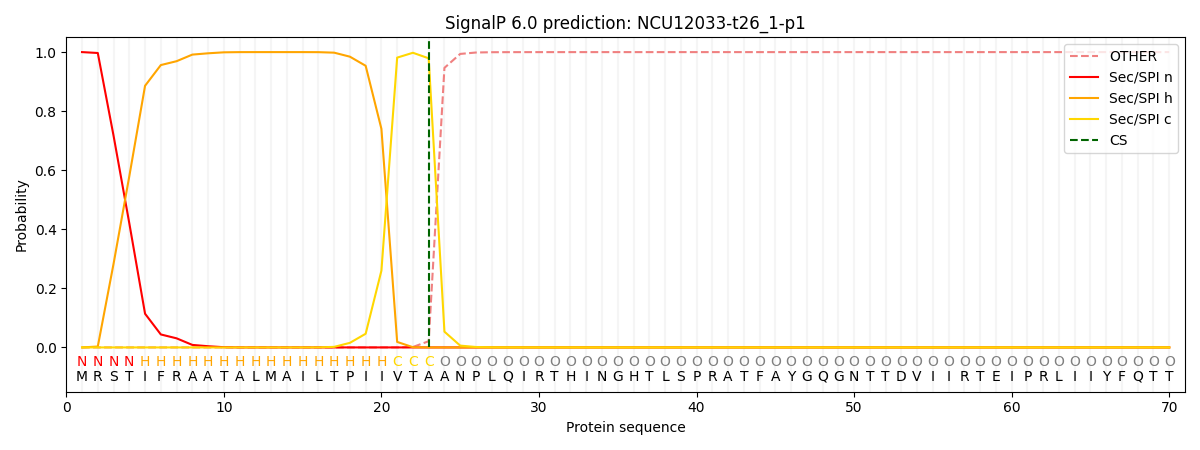
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000245 | 0.999739 | CS pos: 23-24. Pr: 0.9789 |