You are browsing environment: FUNGIDB
CAZyme Information: MELLADRAFT_123733-t26_1-p1
You are here: Home > Sequence: MELLADRAFT_123733-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Melampsora larici-populina | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Basidiomycota; Pucciniomycetes; ; Melampsoraceae; Melampsora; Melampsora larici-populina | |||||||||||
| CAZyme ID | MELLADRAFT_123733-t26_1-p1 | |||||||||||
| CAZy Family | CE4 | |||||||||||
| CAZyme Description | cutinase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE5 | 66 | 253 | 1e-31 | 0.9947089947089947 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 395860 | Cutinase | 1.72e-23 | 66 | 253 | 1 | 173 | Cutinase. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.84e-21 | 65 | 254 | 31 | 211 | |
| 3.06e-21 | 53 | 254 | 27 | 219 | |
| 3.14e-20 | 53 | 254 | 27 | 219 | |
| 9.45e-19 | 65 | 253 | 33 | 211 | |
| 1.16e-18 | 67 | 257 | 60 | 244 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.04e-14 | 66 | 253 | 5 | 201 | A novel cutinase-like protein from Cryptococcus sp. [Cryptococcus sp. S-2],2CZQ_B A novel cutinase-like protein from Cryptococcus sp. [Cryptococcus sp. S-2] |
|
| 1.15e-07 | 65 | 250 | 77 | 244 | Structure of cutinase from Trichoderma reesei in its native form. [Trichoderma reesei QM6a],4PSD_A Structure of Trichoderma reesei cutinase native form. [Trichoderma reesei QM6a],4PSE_A Trichoderma reesei cutinase in complex with a C11Y4 phosphonate inhibitor [Trichoderma reesei QM6a],4PSE_B Trichoderma reesei cutinase in complex with a C11Y4 phosphonate inhibitor [Trichoderma reesei QM6a] |
|
| 1.42e-07 | 66 | 253 | 1 | 206 | Acetylxylan Esterase From P. Purpurogenum Refined At 1.10 Angstroms [Talaromyces purpureogenus] |
|
| 1.42e-07 | 66 | 253 | 1 | 206 | Iodinated Complex Of Acetyl Xylan Esterase At 1.80 Angstroms [Talaromyces purpureogenus] |
|
| 4.82e-07 | 67 | 253 | 2 | 206 | ACETYLXYLAN ESTERASE AT 0.90 ANGSTROM RESOLUTION [Talaromyces purpureogenus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.92e-15 | 59 | 253 | 27 | 213 | Carboxylesterase Culp1 OS=Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv) OX=83332 GN=cut7 PE=1 SV=1 |
|
| 4.92e-15 | 59 | 253 | 27 | 213 | Carboxylesterase Culp1 OS=Mycobacterium bovis (strain ATCC BAA-935 / AF2122/97) OX=233413 GN=BQ2027_MB2006C PE=3 SV=1 |
|
| 4.92e-15 | 59 | 253 | 27 | 213 | Carboxylesterase Culp1 OS=Mycobacterium tuberculosis (strain CDC 1551 / Oshkosh) OX=83331 GN=MT2037 PE=3 SV=1 |
|
| 3.96e-14 | 67 | 253 | 47 | 226 | Phospholipase Culp4 OS=Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv) OX=83332 GN=cut4 PE=1 SV=3 |
|
| 6.65e-09 | 65 | 255 | 42 | 228 | Probable carboxylesterase Culp3 OS=Mycobacterium bovis (strain ATCC BAA-935 / AF2122/97) OX=233413 GN=cut3 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
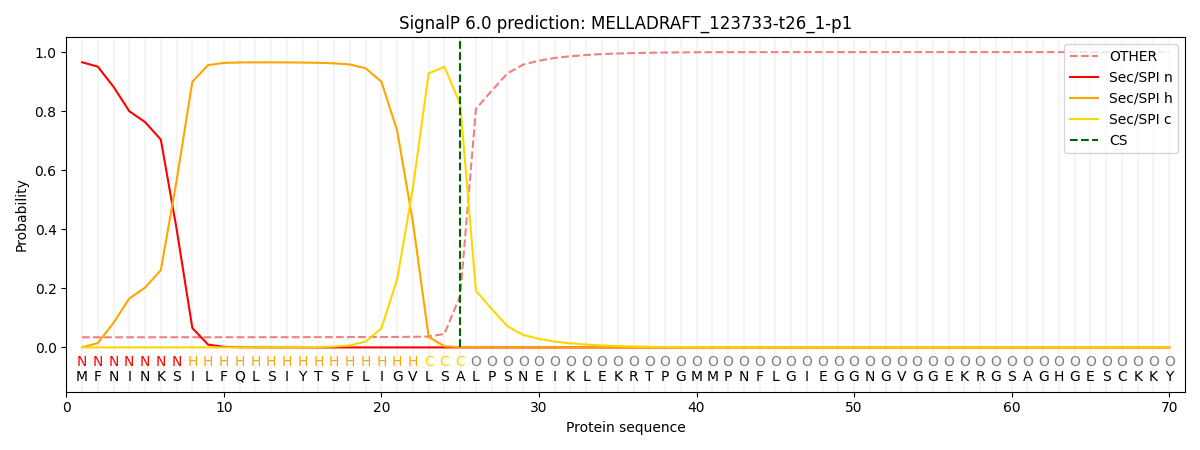
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.041053 | 0.958909 | CS pos: 25-26. Pr: 0.8237 |
