You are browsing environment: FUNGIDB
CAZyme Information: KRZ99961.1
You are here: Home > Sequence: KRZ99961.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Debaryomyces fabryi | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Saccharomycetes; ; Debaryomycetaceae; Debaryomyces; Debaryomyces fabryi | |||||||||||
| CAZyme ID | KRZ99961.1 | |||||||||||
| CAZy Family | GH17 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 2.4.1.-:29 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT24 | 1218 | 1466 | 2e-129 | 0.9959677419354839 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 408203 | Glyco_transf_24 | 0.0 | 1218 | 1486 | 1 | 268 | Glucosyltransferase 24. This is the catalytic domain found in UDP-glucose:glycoprotein glucosyltransferase (UGGT). This domain belongs to glucosyltransferase 24 family (GT24) A-type domain. The GT domain displays the expected glycosyltransferase type A (GT-A) fold. |
| 133054 | GT8_HUGT1_C_like | 1.61e-161 | 1218 | 1466 | 1 | 248 | The C-terminal domain of HUGT1-like is highly homologous to the GT 8 family. C-terminal domain of glycoprotein glucosyltransferase (UGT). UGT is a large glycoprotein whose C-terminus contains the catalytic activity. This catalytic C-terminal domain is highly homologous to Glycosyltransferase Family 8 (GT 8) and contains the DXD motif that coordinates donor sugar binding, characteristic for Family 8 glycosyltransferases. GT 8 proteins are retaining enzymes based on the relative anomeric stereochemistry of the substrate and product in the reaction catalyzed. The non-catalytic N-terminal portion of the human UTG1 (HUGT1) has been shown to monitor the protein folding status and activate its glucosyltransferase activity. |
| 132996 | Glyco_transf_8 | 6.82e-55 | 1218 | 1459 | 1 | 241 | Members of glycosyltransferase family 8 (GT-8) are involved in lipopolysaccharide biosynthesis and glycogen synthesis. Members of this family are involved in lipopolysaccharide biosynthesis and glycogen synthesis. GT-8 comprises enzymes with a number of known activities: lipopolysaccharide galactosyltransferase, lipopolysaccharide glucosyltransferase 1, glycogenin glucosyltransferase, and N-acetylglucosaminyltransferase. GT-8 enzymes contains a conserved DXD motif which is essential in the coordination of a catalytic divalent cation, most commonly Mn2+. |
| 408201 | Thioredoxin_14 | 5.14e-37 | 478 | 696 | 10 | 248 | Thioredoxin-like domain. This is the third out of four TRXL(thioredoxin-like) domains found in UDP-glucose:glycoprotein glucosyltransferase (UGGT). |
| 408199 | Thioredoxin_12 | 2.11e-34 | 42 | 257 | 1 | 185 | Thioredoxin-like domain. This is one of four TRXL(thioredoxin-like) domains found in UDP-glucose:glycoprotein glucosyltransferase (UGGT). |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 1528 | 1 | 1532 | |
| 0.0 | 12 | 1487 | 7 | 1475 | |
| 0.0 | 30 | 1528 | 26 | 1447 | |
| 0.0 | 30 | 1498 | 26 | 1418 | |
| 0.0 | 2 | 1528 | 10 | 1500 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.18e-202 | 34 | 1492 | 13 | 1453 | Chain A, UDP-glucose-glycoprotein glucosyltransferase-like protein [Thermochaetoides thermophila DSM 1495],5MZO_A Chain A, UDP-glucose-glycoprotein glucosyltransferase-like protein [Thermochaetoides thermophila DSM 1495],5N2J_A Chain A, UDP-glucose-glycoprotein glucosyltransferase-like protein [Thermochaetoides thermophila DSM 1495],5N2J_B Chain B, UDP-glucose-glycoprotein glucosyltransferase-like protein [Thermochaetoides thermophila DSM 1495],6TRF_A Chain A, UDP-glucose-glycoprotein glucosyltransferase-like protein [Thermochaetoides thermophila DSM 1495] |
|
| 1.18e-202 | 34 | 1492 | 13 | 1453 | Chain A, UDP-glucose-glycoprotein glucosyltransferase-like protein [Thermochaetoides thermophila DSM 1495] |
|
| 4.46e-202 | 34 | 1492 | 13 | 1453 | Chain A, UDP-glucose-glycoprotein glucosyltransferase-like protein [Thermochaetoides thermophila DSM 1495] |
|
| 6.66e-173 | 34 | 1492 | 13 | 1219 | Chain A, UDP-glucose-glycoprotein glucosyltransferase-like protein,UDP-glucose-glycoprotein glucosyltransferase-like protein [Thermochaetoides thermophila DSM 1495],6TS2_B Chain B, UDP-glucose-glycoprotein glucosyltransferase-like protein,UDP-glucose-glycoprotein glucosyltransferase-like protein [Thermochaetoides thermophila DSM 1495],6TS2_C Chain C, UDP-glucose-glycoprotein glucosyltransferase-like protein,UDP-glucose-glycoprotein glucosyltransferase-like protein [Thermochaetoides thermophila DSM 1495],6TS2_D Chain D, UDP-glucose-glycoprotein glucosyltransferase-like protein,UDP-glucose-glycoprotein glucosyltransferase-like protein [Thermochaetoides thermophila DSM 1495] |
|
| 5.43e-168 | 34 | 1428 | 6 | 1382 | Chain A, UDP-glucose-glycoprotein glucosyltransferase-like protein [Thermochaetoides thermophila DSM 1495],6TS8_B Chain B, UDP-glucose-glycoprotein glucosyltransferase-like protein [Thermochaetoides thermophila DSM 1495] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.88e-195 | 98 | 1506 | 56 | 1444 | UDP-glucose:glycoprotein glucosyltransferase OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=gpt1 PE=1 SV=2 |
|
| 2.52e-161 | 36 | 1495 | 34 | 1516 | UDP-glucose:glycoprotein glucosyltransferase OS=Drosophila melanogaster OX=7227 GN=Uggt PE=1 SV=2 |
|
| 1.06e-154 | 165 | 1511 | 133 | 1516 | UDP-glucose:glycoprotein glucosyltransferase 2 OS=Homo sapiens OX=9606 GN=UGGT2 PE=1 SV=4 |
|
| 1.67e-151 | 204 | 1486 | 185 | 1523 | UDP-glucose:glycoprotein glucosyltransferase 1 OS=Mus musculus OX=10090 GN=Uggt1 PE=1 SV=4 |
|
| 8.33e-150 | 176 | 1486 | 156 | 1523 | UDP-glucose:glycoprotein glucosyltransferase 1 OS=Homo sapiens OX=9606 GN=UGGT1 PE=1 SV=3 |
SignalP and Lipop Annotations help
This protein is predicted as SP
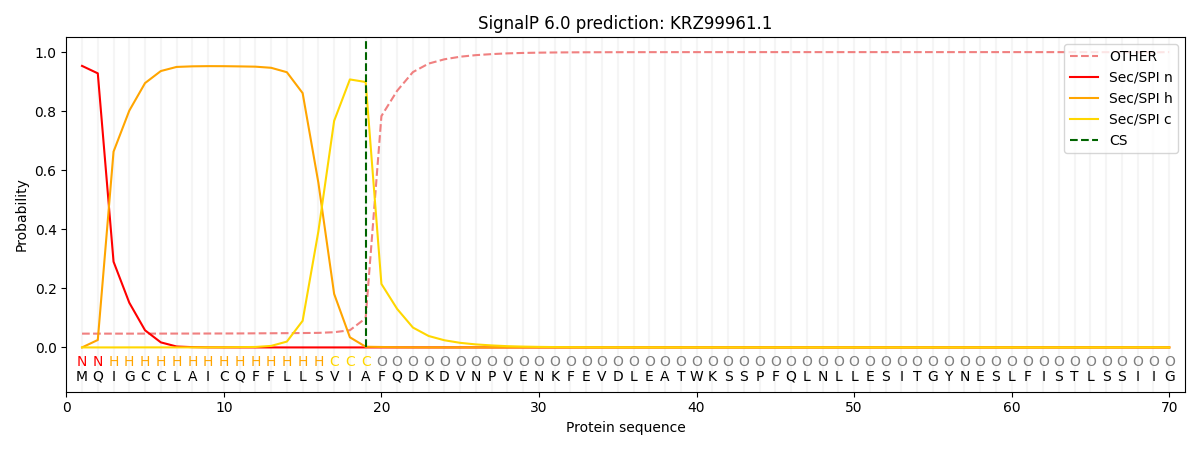
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.053466 | 0.946487 | CS pos: 19-20. Pr: 0.8987 |
