You are browsing environment: FUNGIDB
CAZyme Information: KNZ54879.1
You are here: Home > Sequence: KNZ54879.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
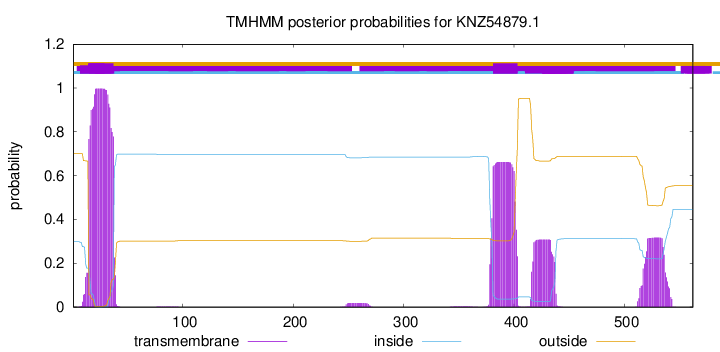
TMHMM annotations
Basic Information help
| Species | Puccinia sorghi | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Basidiomycota; Pucciniomycetes; ; Pucciniaceae; Puccinia; Puccinia sorghi | |||||||||||
| CAZyme ID | KNZ54879.1 | |||||||||||
| CAZy Family | GH26 | |||||||||||
| CAZyme Description | Ceramide glucosyltransferase [Source:UniProtKB/TrEMBL;Acc:A0A0L6V2B2] | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 2.4.1.80:3 | 2.4.1.-:1 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT21 | 80 | 376 | 7.1e-74 | 0.9957081545064378 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 133012 | Glucosylceramide_synthase | 2.15e-59 | 80 | 376 | 1 | 196 | Glucosylceramide synthase catalyzes the first glycosylation step of glycosphingolipid synthesis. UDP-glucose:N-acylsphingosine D-glucosyltransferase (glucosylceramide synthase or ceramide glucosyltransferase) catalyzes the first glycosylation step of glycosphingolipid synthesis. Its product, glucosylceramide, serves as the core of more than 300 glycosphingolipids (GSL). GSLs are a group of membrane components that have the lipid portion embedded in the outer plasma membrane leaflet and the sugar chains extended to the outer environment. Several lines of evidence suggest the importance of GSLs in various cellular processes such as differentiation, adhesion, proliferation, and cell-cell recognition. In pathogenic fungus Cryptococcus neoformans, glucosylceramide serves as an antigen that elicits an antibody response in patients and it is essential for fungal growth in host extracellular environment. |
| 404400 | Glyco_transf_21 | 7.21e-27 | 137 | 374 | 1 | 174 | Glycosyl transferase family 21. This is a family of ceramide beta-glucosyltransferases - EC:2.4.1.80. |
| 224136 | BcsA | 1.19e-09 | 61 | 443 | 35 | 348 | Glycosyltransferase, catalytic subunit of cellulose synthase and poly-beta-1,6-N-acetylglucosamine synthase [Cell motility]. |
| 404520 | Glyco_tranf_2_3 | 2.77e-07 | 95 | 375 | 17 | 230 | Glycosyltransferase like family 2. Members of this family of prokaryotic proteins include putative glucosyltransferase, which are involved in bacterial capsule biosynthesis. |
| 133016 | Succinoglycan_BP_ExoA | 0.002 | 95 | 207 | 15 | 124 | ExoA is involved in the biosynthesis of succinoglycan. Succinoglycan Biosynthesis Protein ExoA catalyzes the formation of a beta-1,3 linkage of the second sugar (glucose) of the succinoglycan with the galactose on the lipid carrie. Succinoglycan is an acidic exopolysaccharide that is important for invasion of the nodules. Succinoglycan is a high-molecular-weight polymer composed of repeating octasaccharide units. These units are synthesized on membrane-bound isoprenoid lipid carriers, beginning with galactose followed by seven glucose molecules, and modified by the addition of acetate, succinate, and pyruvate. ExoA is a membrane protein with a transmembrance domain at c-terminus. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 6.26e-80 | 80 | 558 | 51 | 430 | |
| 3.71e-73 | 77 | 436 | 49 | 374 | |
| 5.20e-73 | 16 | 446 | 21 | 400 | |
| 5.20e-73 | 16 | 446 | 21 | 400 | |
| 2.94e-72 | 70 | 547 | 34 | 405 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.45e-60 | 65 | 544 | 40 | 431 | Ceramide glucosyltransferase OS=Cryptococcus neoformans var. grubii serotype A (strain H99 / ATCC 208821 / CBS 10515 / FGSC 9487) OX=235443 GN=GCS1 PE=1 SV=1 |
|
| 8.04e-54 | 13 | 437 | 40 | 429 | Ceramide glucosyltransferase OS=Komagataella phaffii (strain GS115 / ATCC 20864) OX=644223 GN=PAS_chr3_0357 PE=1 SV=1 |
|
| 2.89e-51 | 21 | 544 | 18 | 529 | Ceramide glucosyltransferase OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=HSX11 PE=1 SV=1 |
|
| 1.18e-39 | 12 | 393 | 4 | 351 | Ceramide glucosyltransferase OS=Magnaporthe oryzae (strain 70-15 / ATCC MYA-4617 / FGSC 8958) OX=242507 GN=MGG_10668 PE=1 SV=1 |
|
| 1.73e-38 | 71 | 446 | 42 | 347 | Ceramide glucosyltransferase-B OS=Xenopus laevis OX=8355 GN=ugcg-b PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER

| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000066 | 0.000000 |