You are browsing environment: FUNGIDB
CAZyme Information: KLU86697.1
You are here: Home > Sequence: KLU86697.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Magnaporthiopsis poae | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Magnaporthaceae; Magnaporthiopsis; Magnaporthiopsis poae | |||||||||||
| CAZyme ID | KLU86697.1 | |||||||||||
| CAZy Family | GH16 | |||||||||||
| CAZyme Description | xyloglucanase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.151:27 | 3.2.1.150:4 | - |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH74 | 77 | 185 | 7.7e-23 | 0.45064377682403434 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 406353 | Sortilin-Vps10 | 3.26e-06 | 57 | 275 | 3 | 217 | Sortilin, neurotensin receptor 3,. Sortilin, also known in mammals as neurotensin receptor-3, is the archetypical member of a Vps10-domain (Vps10-D) that binds neurotrophic factors and neuropeptides. This domain constitutes the entire luminal part of Sortilin and is activated in the trans-Golgi network by enzymatic propeptide cleavage. The structure of the domain has been determined as a ten-bladed propeller, with up to 9 BNR or beta-hairpin turns in it. The mature receptor binds various ligands, including its own propeptide (Sort-pro), neurotensin, the pro-forms of nerve growth factor-beta (NGF)6 and brain-derived neurotrophic factor (BDNF)7, lipoprotein lipase (LpL), apo lipoprotein AV14 and the receptor-associated protein (RAP)1. |
| 271234 | Sialidase_non-viral | 7.45e-05 | 104 | 252 | 165 | 296 | Non-viral sialidases. Sialidases or neuraminidases function to bind and hydrolyze terminal sialic acid residues from various glycoconjugates, they play vital roles in pathogenesis, bacterial nutrition and cellular interactions. They have a six-bladed, beta-propeller fold with the non-viral sialidases containing 2-5 Asp-box motifs (most commonly Ser/Thr-X-Asp-[X]-Gly-X-Thr- Trp/Phe). This CD includes eubacterial and eukaryotic sialidases. |
| 406353 | Sortilin-Vps10 | 0.003 | 113 | 252 | 1 | 112 | Sortilin, neurotensin receptor 3,. Sortilin, also known in mammals as neurotensin receptor-3, is the archetypical member of a Vps10-domain (Vps10-D) that binds neurotrophic factors and neuropeptides. This domain constitutes the entire luminal part of Sortilin and is activated in the trans-Golgi network by enzymatic propeptide cleavage. The structure of the domain has been determined as a ten-bladed propeller, with up to 9 BNR or beta-hairpin turns in it. The mature receptor binds various ligands, including its own propeptide (Sort-pro), neurotensin, the pro-forms of nerve growth factor-beta (NGF)6 and brain-derived neurotrophic factor (BDNF)7, lipoprotein lipase (LpL), apo lipoprotein AV14 and the receptor-associated protein (RAP)1. |
| 411995 | MALA | 0.003 | 284 | 348 | 185 | 248 | Mala s 1 allergenic protein and similar proteins. This family includes the yeast Malassezia sympodialis allergen Mala s 1 which is localized in the cell wall and exposed on the cell surface. It can elicit specific IgE and T-cell activity in patients with atopic eczema (AE), a chronic inflammatory disease. Mala s 1 does not show any significant sequence homology to characterized proteins. However, its structure is a beta-propeller which is a novel fold among allergens. |
| 214740 | VPS10 | 0.004 | 113 | 266 | 55 | 215 | VPS10 domain. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.55e-277 | 9 | 506 | 10 | 505 | |
| 7.28e-277 | 9 | 506 | 10 | 503 | |
| 8.90e-275 | 1 | 506 | 1 | 512 | |
| 8.90e-275 | 1 | 506 | 1 | 512 | |
| 1.22e-272 | 1 | 506 | 1 | 512 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.47e-174 | 21 | 506 | 6 | 485 | Crystal structure of the catalytic domain from GH74 enzyme PoGH74 from Paenibacillus odorifer, D60A mutant in complex with XXLG and XGXXLG xyloglucan [Paenibacillus odorifer] |
|
| 7.24e-173 | 20 | 506 | 2 | 484 | Crystal structure of Streptomyces rapamycinicus GH74 in complex with xyloglucan fragments XLLG and XXXG [Streptomyces rapamycinicus NRRL 5491],6P2O_B Crystal structure of Streptomyces rapamycinicus GH74 in complex with xyloglucan fragments XLLG and XXXG [Streptomyces rapamycinicus NRRL 5491] |
|
| 7.55e-172 | 21 | 506 | 6 | 485 | Crystal structure of the catalytic domain from GH74 enzyme PoGH74 from Paenibacillus odorifer, apoenzyme [Paenibacillus odorifer],6MGJ_B Crystal structure of the catalytic domain from GH74 enzyme PoGH74 from Paenibacillus odorifer, apoenzyme [Paenibacillus odorifer],6MGJ_C Crystal structure of the catalytic domain from GH74 enzyme PoGH74 from Paenibacillus odorifer, apoenzyme [Paenibacillus odorifer],6MGJ_D Crystal structure of the catalytic domain from GH74 enzyme PoGH74 from Paenibacillus odorifer, apoenzyme [Paenibacillus odorifer],6MGJ_E Crystal structure of the catalytic domain from GH74 enzyme PoGH74 from Paenibacillus odorifer, apoenzyme [Paenibacillus odorifer],6MGJ_F Crystal structure of the catalytic domain from GH74 enzyme PoGH74 from Paenibacillus odorifer, apoenzyme [Paenibacillus odorifer],6MGJ_G Crystal structure of the catalytic domain from GH74 enzyme PoGH74 from Paenibacillus odorifer, apoenzyme [Paenibacillus odorifer],6MGJ_H Crystal structure of the catalytic domain from GH74 enzyme PoGH74 from Paenibacillus odorifer, apoenzyme [Paenibacillus odorifer],6MGK_A Crystal structure of the catalytic domain from GH74 enzyme PoGH74 from Paenibacillus odorifer, in complex with XLX xyloglucan [Paenibacillus odorifer],6MGK_B Crystal structure of the catalytic domain from GH74 enzyme PoGH74 from Paenibacillus odorifer, in complex with XLX xyloglucan [Paenibacillus odorifer],6MGK_C Crystal structure of the catalytic domain from GH74 enzyme PoGH74 from Paenibacillus odorifer, in complex with XLX xyloglucan [Paenibacillus odorifer],6MGK_D Crystal structure of the catalytic domain from GH74 enzyme PoGH74 from Paenibacillus odorifer, in complex with XLX xyloglucan [Paenibacillus odorifer] |
|
| 3.52e-170 | 20 | 506 | 5 | 486 | Crystal structure of Paenibacillus graminis GH74 (PgGH74) [Paenibacillus graminis] |
|
| 8.68e-168 | 21 | 506 | 3 | 484 | Structure of a Putative Xyloglucanase from the Cellulolytic Bacteria Streptomyces sp. SirexAA-E [Streptomyces sp. SirexAA-E],5JWZ_B Structure of a Putative Xyloglucanase from the Cellulolytic Bacteria Streptomyces sp. SirexAA-E [Streptomyces sp. SirexAA-E] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.58e-275 | 1 | 506 | 1 | 512 | Xyloglucanase OS=Hypocrea jecorina (strain QM6a) OX=431241 GN=cel74a PE=1 SV=1 |
|
| 1.04e-170 | 21 | 506 | 38 | 519 | Xyloglucanase OS=Paenibacillus sp. OX=58172 GN=xeg74 PE=1 SV=1 |
|
| 1.49e-149 | 15 | 501 | 32 | 514 | Xyloglucanase Xgh74A OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=xghA PE=3 SV=1 |
|
| 5.92e-149 | 15 | 501 | 32 | 514 | Xyloglucanase Xgh74A OS=Acetivibrio thermocellus OX=1515 GN=xghA PE=1 SV=1 |
|
| 1.85e-74 | 12 | 507 | 26 | 538 | Oligoxyloglucan-reducing end-specific xyloglucanase OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=xgcA PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
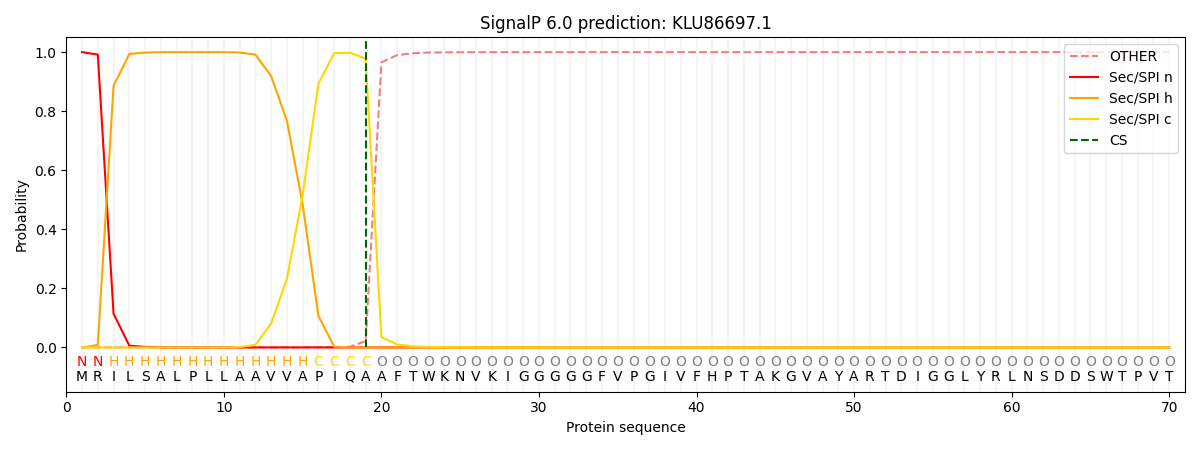
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000223 | 0.999714 | CS pos: 19-20. Pr: 0.9783 |
