You are browsing environment: FUNGIDB
CAZyme Information: KIX08809.1
You are here: Home > Sequence: KIX08809.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Rhinocladiella mackenziei | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Herpotrichiellaceae; Rhinocladiella; Rhinocladiella mackenziei | |||||||||||
| CAZyme ID | KIX08809.1 | |||||||||||
| CAZy Family | GT21 | |||||||||||
| CAZyme Description | GMC_OxRdtase_N domain-containing protein [Source:UniProtKB/TrEMBL;Acc:A0A0D2IS29] | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 1.1.3.13:12 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| AA3 | 189 | 559 | 9.2e-147 | 0.6288135593220339 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 225186 | BetA | 9.00e-62 | 13 | 552 | 7 | 531 | Choline dehydrogenase or related flavoprotein [Lipid transport and metabolism, General function prediction only]. |
| 235000 | PRK02106 | 2.37e-56 | 12 | 553 | 4 | 530 | choline dehydrogenase; Validated |
| 398739 | GMC_oxred_C | 2.33e-30 | 389 | 550 | 1 | 143 | GMC oxidoreductase. This domain found associated with pfam00732. |
| 366272 | GMC_oxred_N | 3.95e-16 | 100 | 279 | 30 | 216 | GMC oxidoreductase. This family of proteins bind FAD as a cofactor. |
| 215420 | PLN02785 | 3.63e-10 | 12 | 547 | 54 | 569 | Protein HOTHEAD |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.51e-210 | 1 | 560 | 1 | 604 | |
| 4.58e-208 | 1 | 558 | 1 | 603 | |
| 4.37e-199 | 1 | 559 | 1 | 601 | |
| 4.34e-197 | 1 | 562 | 1 | 605 | |
| 1.28e-196 | 1 | 560 | 1 | 604 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.00e-126 | 12 | 560 | 5 | 626 | Chain A, Alcohol oxidase [Phanerodontia chrysosporium],6H3O_B Chain B, Alcohol oxidase [Phanerodontia chrysosporium],6H3O_C Chain C, Alcohol oxidase [Phanerodontia chrysosporium],6H3O_D Chain D, Alcohol oxidase [Phanerodontia chrysosporium],6H3O_E Chain E, Alcohol oxidase [Phanerodontia chrysosporium],6H3O_F Chain F, Alcohol oxidase [Phanerodontia chrysosporium],6H3O_G Chain G, Alcohol oxidase [Phanerodontia chrysosporium],6H3O_H Chain H, Alcohol oxidase [Phanerodontia chrysosporium] |
|
| 5.62e-126 | 12 | 560 | 5 | 626 | Chain A, Alcohol oxidase [Phanerodontia chrysosporium],6H3G_B Chain B, Alcohol oxidase [Phanerodontia chrysosporium],6H3G_C Chain C, Alcohol oxidase [Phanerodontia chrysosporium],6H3G_D Chain D, Alcohol oxidase [Phanerodontia chrysosporium],6H3G_E Chain E, Alcohol oxidase [Phanerodontia chrysosporium],6H3G_F Chain F, Alcohol oxidase [Phanerodontia chrysosporium],6H3G_G Chain G, Alcohol oxidase [Phanerodontia chrysosporium],6H3G_H Chain H, Alcohol oxidase [Phanerodontia chrysosporium] |
|
| 1.09e-103 | 12 | 559 | 5 | 637 | Alcohol Oxidase AOX1 from Pichia Pastoris [Komagataella phaffii CBS 7435],5HSA_B Alcohol Oxidase AOX1 from Pichia Pastoris [Komagataella phaffii CBS 7435],5HSA_C Alcohol Oxidase AOX1 from Pichia Pastoris [Komagataella phaffii CBS 7435],5HSA_D Alcohol Oxidase AOX1 from Pichia Pastoris [Komagataella phaffii CBS 7435],5HSA_E Alcohol Oxidase AOX1 from Pichia Pastoris [Komagataella phaffii CBS 7435],5HSA_F Alcohol Oxidase AOX1 from Pichia Pastoris [Komagataella phaffii CBS 7435],5HSA_G Alcohol Oxidase AOX1 from Pichia Pastoris [Komagataella phaffii CBS 7435],5HSA_H Alcohol Oxidase AOX1 from Pichia Pastoris [Komagataella phaffii CBS 7435],5I68_A Chain A, Alcohol oxidase 1 [Komagataella pastoris] |
|
| 3.21e-45 | 17 | 557 | 9 | 585 | Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE2_B Chain B, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE3_A Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE4_A Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE4_B Chain B, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE5_A Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE5_B Chain B, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE6_A Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE6_B Chain B, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE7_A Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE7_B Chain B, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],7AA2_A Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],7AA2_B Chain B, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495] |
|
| 2.12e-43 | 8 | 551 | 2 | 557 | Chain AAA, Fatty acid Photodecarboxylase [Chlorella variabilis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.81e-113 | 7 | 559 | 3 | 638 | Alcohol oxidase OS=Pichia angusta OX=870730 GN=MOX PE=1 SV=1 |
|
| 9.27e-113 | 12 | 559 | 5 | 637 | Alcohol oxidase OS=Candida boidinii OX=5477 GN=AOD1 PE=1 SV=1 |
|
| 2.85e-103 | 12 | 559 | 5 | 637 | Alcohol oxidase 2 OS=Komagataella phaffii (strain ATCC 76273 / CBS 7435 / CECT 11047 / NRRL Y-11430 / Wegner 21-1) OX=981350 GN=AOX2 PE=2 SV=1 |
|
| 2.85e-103 | 12 | 559 | 5 | 637 | Alcohol oxidase 2 OS=Komagataella phaffii (strain GS115 / ATCC 20864) OX=644223 GN=AOX2 PE=3 SV=1 |
|
| 5.62e-103 | 12 | 559 | 5 | 637 | Alcohol oxidase 1 OS=Komagataella phaffii (strain ATCC 76273 / CBS 7435 / CECT 11047 / NRRL Y-11430 / Wegner 21-1) OX=981350 GN=AOX1 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
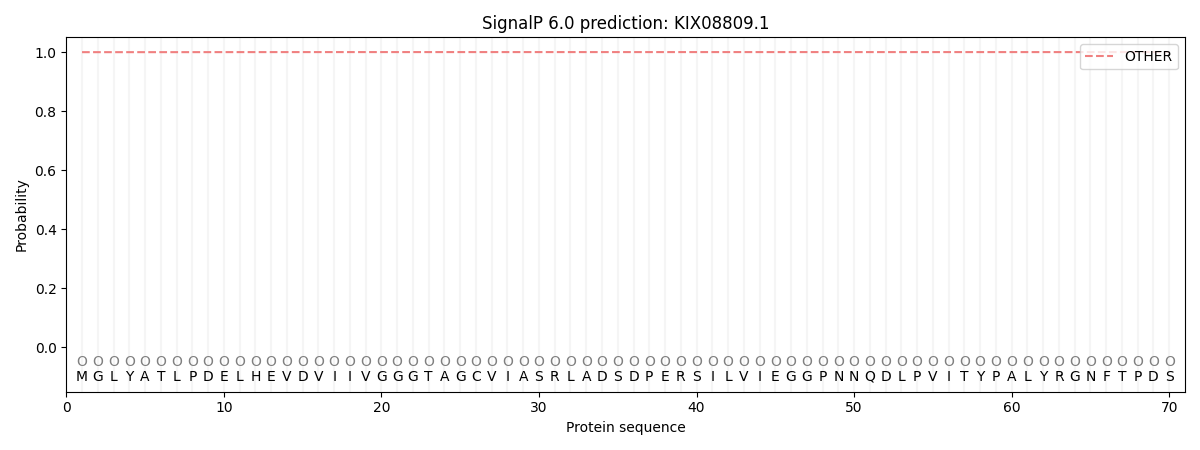
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.999869 | 0.000175 |
