You are browsing environment: FUNGIDB
CAZyme Information: KIW96593.1
You are here: Home > Sequence: KIW96593.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
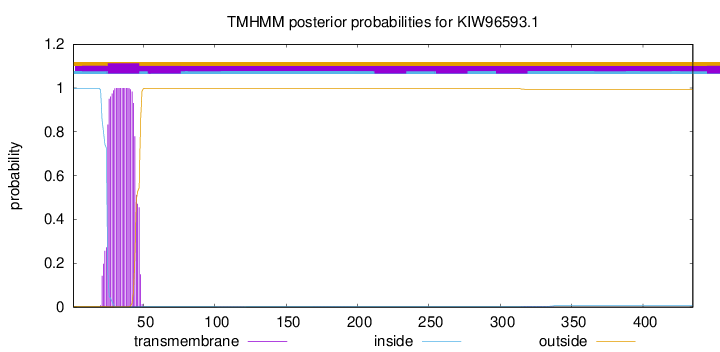
TMHMM annotations
Basic Information help
| Species | Cladophialophora bantiana | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Herpotrichiellaceae; Cladophialophora; Cladophialophora bantiana | |||||||||||
| CAZyme ID | KIW96593.1 | |||||||||||
| CAZy Family | GT2 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 92 | 403 | 1.1e-141 | 0.9904761904761905 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 395098 | Cellulase | 5.62e-16 | 93 | 399 | 1 | 270 | Cellulase (glycosyl hydrolase family 5). |
| 225344 | BglC | 1.78e-09 | 132 | 435 | 82 | 399 | Aryl-phospho-beta-D-glucosidase BglC, GH1 family [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 3.17e-156 | 18 | 435 | 80 | 505 | |
| 8.76e-155 | 28 | 432 | 87 | 497 | |
| 1.03e-154 | 46 | 435 | 112 | 505 | |
| 1.24e-154 | 28 | 432 | 87 | 497 | |
| 1.38e-154 | 61 | 432 | 210 | 586 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 9.55e-45 | 87 | 419 | 8 | 338 | Acidothermus Cellulolyticus Endocellulase E1 Catalytic Domain In Complex With A Cellotetraose [Acidothermus cellulolyticus],1ECE_B Acidothermus Cellulolyticus Endocellulase E1 Catalytic Domain In Complex With A Cellotetraose [Acidothermus cellulolyticus] |
|
| 1.86e-44 | 87 | 419 | 8 | 338 | Chain A, ENDOCELLULASE E1 FROM A. CELLULOLYTICUS [Acidothermus cellulolyticus],1VRX_B Chain B, ENDOCELLULASE E1 FROM A. CELLULOLYTICUS [Acidothermus cellulolyticus] |
|
| 1.96e-27 | 90 | 423 | 7 | 335 | Catalytic domain of the major endoglucanase from Xanthomonas campestris pv. campestris [Xanthomonas campestris pv. campestris str. ATCC 33913],4TUF_B Catalytic domain of the major endoglucanase from Xanthomonas campestris pv. campestris [Xanthomonas campestris pv. campestris str. ATCC 33913],4TUF_C Catalytic domain of the major endoglucanase from Xanthomonas campestris pv. campestris [Xanthomonas campestris pv. campestris str. ATCC 33913],4TUF_D Catalytic domain of the major endoglucanase from Xanthomonas campestris pv. campestris [Xanthomonas campestris pv. campestris str. ATCC 33913] |
|
| 2.09e-26 | 91 | 419 | 19 | 363 | Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3],3W6M_B Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3],3W6M_C Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3],4DM2_A Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3],4DM2_B Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3],4DM2_C Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3] |
|
| 3.91e-26 | 91 | 419 | 19 | 363 | The Crystal Structure of Cellulase-Inhibitor Complex. [Pyrococcus horikoshii OT3],3VVG_B The Crystal Structure of Cellulase-Inhibitor Complex. [Pyrococcus horikoshii OT3],3VVG_C The Crystal Structure of Cellulase-Inhibitor Complex. [Pyrococcus horikoshii OT3],3W6L_A Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3],3W6L_B Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3],3W6L_C Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3],4DM1_A Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3],4DM1_B Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3],4DM1_C Contribution of disulfide bond toward thermostability in hyperthermostable endocellulase [Pyrococcus horikoshii OT3] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.98e-42 | 87 | 419 | 49 | 379 | Endoglucanase E1 OS=Acidothermus cellulolyticus (strain ATCC 43068 / DSM 8971 / 11B) OX=351607 GN=Acel_0614 PE=1 SV=1 |
|
| 1.54e-35 | 87 | 419 | 41 | 369 | Endoglucanase OS=Paenibacillus polymyxa OX=1406 PE=3 SV=2 |
|
| 2.40e-31 | 84 | 417 | 30 | 379 | Glycosyl hydrolase 5 family protein OS=Chamaecyparis obtusa OX=13415 PE=1 SV=1 |
|
| 9.94e-27 | 85 | 419 | 46 | 400 | Endoglucanase D OS=Cellulomonas fimi OX=1708 GN=cenD PE=3 SV=1 |
|
| 5.45e-26 | 90 | 423 | 32 | 360 | Major extracellular endoglucanase OS=Xanthomonas campestris pv. campestris (strain ATCC 33913 / DSM 3586 / NCPPB 528 / LMG 568 / P 25) OX=190485 GN=engXCA PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
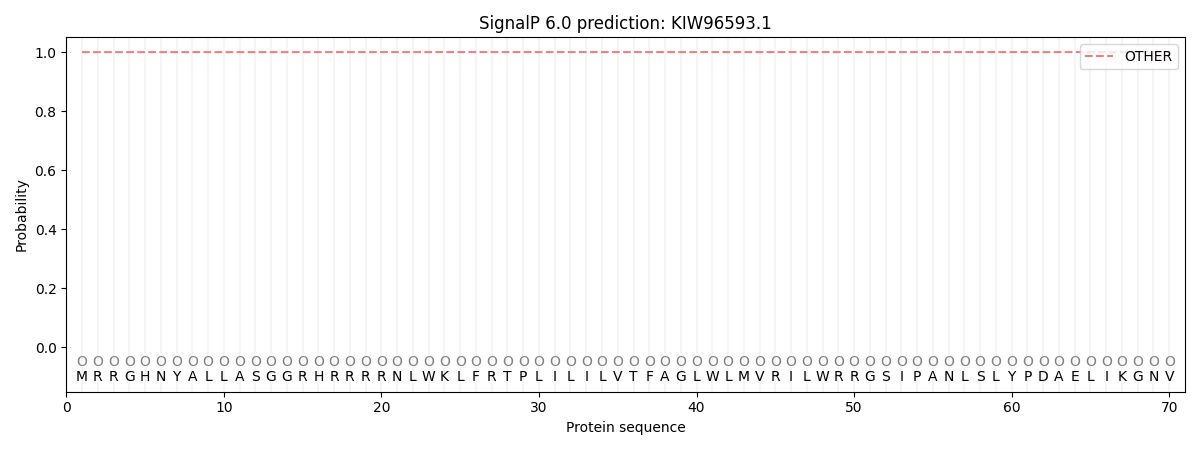
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000018 | 0.000032 |