You are browsing environment: FUNGIDB
CAZyme Information: KIW93065.1
You are here: Home > Sequence: KIW93065.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Cladophialophora bantiana | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Herpotrichiellaceae; Cladophialophora; Cladophialophora bantiana | |||||||||||
| CAZyme ID | KIW93065.1 | |||||||||||
| CAZy Family | GH28 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 1.10.3.2:4 | 1.10.3.-:1 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| AA1 | 208 | 511 | 5.5e-108 | 0.967948717948718 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 259923 | CuRO_1_MaLCC_like | 1.27e-68 | 216 | 339 | 1 | 122 | The first cupredoxin domain of the fungal laccases similar to Ma-LCC from Melanocarpus albomyces. The subfamily of fungal laccases includes Ma-LCC and similar proteins. Ma-LCC is a multicopper oxidase (MCO) from Melanocarpus albomyces. Its crystal structure contains all four coppers at the mono- and trinuclear copper centers. Laccase is a blue multi-copper enzyme that catalyzes the oxidation of a variety aromatic - notably phenolic and inorganic substances coupled to the reduction of molecular oxygen to water. It has been implicated in a wide spectrum of biological activities and, in particular, plays a key role in morphogenesis, development and lignin metabolism in fungi and plants. Although MCOs have diverse functions, majority of them have three cupredoxin domain repeats that include one mononuclear and one trinuclear copper center. The copper ions are bound in several sites: Type 1, Type 2, and/or Type 3. The ensemble of types 2 and 3 copper is called a trinuclear cluster. MCOs oxidize their substrate by accepting electrons at a mononuclear copper center and transferring them to the active site trinuclear copper center. The cupredoxin domain 1 of 3-domain MCOs contains part the trinuclear copper binding site, which is located at the interface of domains 1 and 3. |
| 259968 | CuRO_3_MaLCC_like | 5.86e-65 | 555 | 708 | 12 | 157 | The third cupredoxin domain of the fungal laccases similar to Ma-LCC from Melanocarpus albomyces. The subfamily of fungal laccases includes Ma-LCC and similar proteins. Ma-LCC is a multicopper oxidase (MCO) from Melanocarpus albomyces. Its crystal structure contains all four coppers at the mono- and trinuclear copper centers. Laccase is a blue multi-copper enzyme that catalyzes the oxidation of a variety aromatic - notably phenolic and inorganic substances coupled to the reduction of molecular oxygen to water. It has been implicated in a wide spectrum of biological activities and, in particular, plays a key role in morphogenesis, development and lignin metabolism in fungi and plants. Although MCOs have diverse functions, majority of them have three cupredoxin domain repeats that include one mononuclear and one trinuclear copper center. The copper ions are bound in several sites: Type 1, Type 2, and/or Type 3. The ensemble of types 2 and 3 copper is called a trinuclear cluster. MCOs oxidize their substrate by accepting electrons at a mononuclear copper center and transferring them to the active site trinuclear copper center. The cupredoxin domain 3 of 3-domain MCOs contains the Type 1 (T1) copper binding site and part the trinuclear copper binding site, which is located at the interface of domains 1 and 3. |
| 225043 | SufI | 1.28e-58 | 214 | 711 | 28 | 450 | Multicopper oxidase with three cupredoxin domains (includes cell division protein FtsP and spore coat protein CotA) [Cell cycle control, cell division, chromosome partitioning, Inorganic ion transport and metabolism, Cell wall/membrane/envelope biogenesis]. |
| 274555 | ascorbase | 7.00e-52 | 238 | 715 | 20 | 532 | L-ascorbate oxidase, plant type. Members of this protein family are the copper-containing enzyme L-ascorbate oxidase (EC 1.10.3.3), also called ascorbase. This family is found in flowering plants, and shows greater sequence similarity to a family of laccases (EC 1.10.3.2) from plants than to other known ascorbate oxidases. |
| 177843 | PLN02191 | 3.52e-51 | 241 | 714 | 45 | 551 | L-ascorbate oxidase |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 4.87e-204 | 191 | 751 | 224 | 796 | |
| 4.03e-141 | 190 | 734 | 39 | 562 | |
| 2.62e-135 | 189 | 730 | 84 | 611 | |
| 4.67e-132 | 191 | 738 | 44 | 562 | |
| 4.67e-132 | 191 | 738 | 44 | 562 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 9.55e-120 | 178 | 734 | 28 | 567 | Crystal structure of laccase from Botrytis aclada at 1.67 A resolution [Botrytis aclada],4X4K_A Structure of laccase from Botrytis aclada with full copper content [Botrytis aclada] |
|
| 1.89e-119 | 178 | 734 | 28 | 567 | Structure of the L499M mutant of the laccase from B.aclada [Botrytis aclada] |
|
| 4.92e-112 | 190 | 739 | 4 | 551 | Crystal Structure of Laccase from Melanocarpus albomyces in Four Copper Form [Melanocarpus albomyces],1GW0_B Crystal Structure of Laccase from Melanocarpus albomyces in Four Copper Form [Melanocarpus albomyces],2IH8_A A low-dose crystal structure of a recombinant Melanocarpus albomyces laccase [Melanocarpus albomyces],2IH8_B A low-dose crystal structure of a recombinant Melanocarpus albomyces laccase [Melanocarpus albomyces],2IH9_A A high-dose crystal structure of a recombinant Melanocarbus albomyces laccase [Melanocarpus albomyces],2IH9_B A high-dose crystal structure of a recombinant Melanocarbus albomyces laccase [Melanocarpus albomyces],3FU7_B Melanocarpus albomyces laccase crystal soaked (4 sec) with 2,6-dimethoxyphenol [Melanocarpus albomyces],3FU9_A Melanocarpus albomyces laccase crystal soaked (20 min) with 2,6-dimethoxyphenol [Melanocarpus albomyces],3FU9_B Melanocarpus albomyces laccase crystal soaked (20 min) with 2,6-dimethoxyphenol [Melanocarpus albomyces],3QPK_A Probing oxygen channels in Melanocarpus albomyces laccase [Melanocarpus albomyces],3QPK_B Probing oxygen channels in Melanocarpus albomyces laccase [Melanocarpus albomyces] |
|
| 4.92e-112 | 190 | 739 | 4 | 551 | L559A mutant of Melanocarpus albomyces laccase [Melanocarpus albomyces],3DKH_B L559A mutant of Melanocarpus albomyces laccase [Melanocarpus albomyces] |
|
| 6.91e-112 | 190 | 739 | 4 | 551 | Near-atomic resolution structure of a Melanocarpus albomyces laccase [Melanocarpus albomyces],2Q9O_B Near-atomic resolution structure of a Melanocarpus albomyces laccase [Melanocarpus albomyces],3FU7_A Melanocarpus albomyces laccase crystal soaked (4 sec) with 2,6-dimethoxyphenol [Melanocarpus albomyces],3FU8_A Melanocarpus albomyces laccase crystal soaked (10 sec) with 2,6-dimethoxyphenol [Melanocarpus albomyces],3FU8_B Melanocarpus albomyces laccase crystal soaked (10 sec) with 2,6-dimethoxyphenol [Melanocarpus albomyces] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.43e-123 | 190 | 753 | 55 | 613 | Laccase OS=Neurospora crassa (strain ATCC 24698 / 74-OR23-1A / CBS 708.71 / DSM 1257 / FGSC 987) OX=367110 GN=lacc PE=1 SV=3 |
|
| 9.33e-123 | 196 | 740 | 51 | 582 | Oxidoreductase OpS5 OS=Beauveria bassiana (strain ARSEF 2860) OX=655819 GN=OpS5 PE=1 SV=1 |
|
| 1.04e-116 | 248 | 707 | 1 | 452 | Laccase-3 (Fragment) OS=Botryotinia fuckeliana OX=40559 GN=lcc3 PE=3 SV=1 |
|
| 2.21e-116 | 190 | 711 | 36 | 552 | Laccase OS=Cryphonectria parasitica OX=5116 GN=LAC-1 PE=3 SV=1 |
|
| 1.94e-114 | 181 | 734 | 31 | 568 | Laccase-2 OS=Botryotinia fuckeliana OX=40559 GN=lcc2 PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
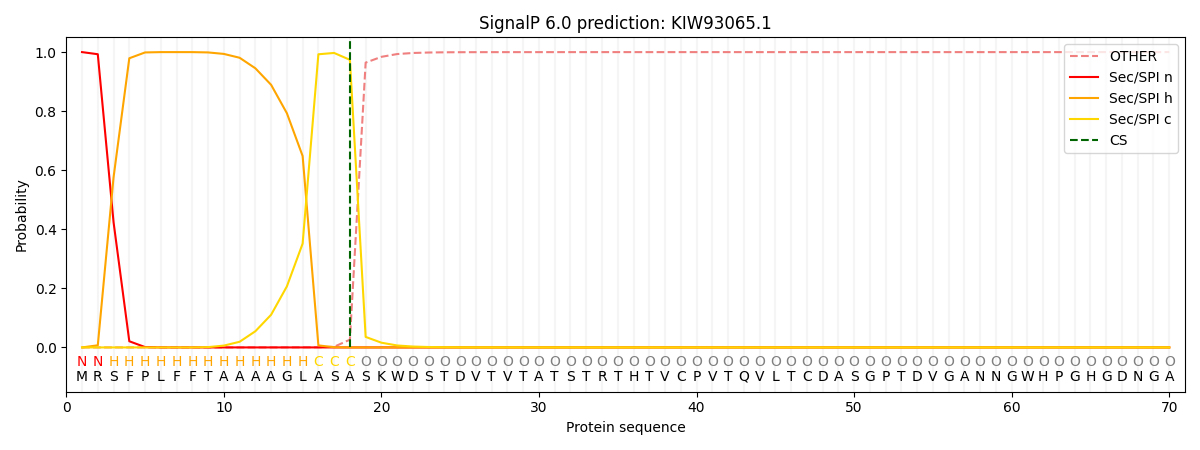
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000249 | 0.999722 | CS pos: 18-19. Pr: 0.9738 |
