You are browsing environment: FUNGIDB
CAZyme Information: KIS67886.1
You are here: Home > Sequence: KIS67886.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Ustilago maydis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Basidiomycota; Ustilaginomycetes; ; Ustilaginaceae; Ustilago; Ustilago maydis | |||||||||||
| CAZyme ID | KIS67886.1 | |||||||||||
| CAZy Family | GH16 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 233912; End:238054 Strand: - | |||||||||||
Full Sequence Download help
| MASSLRKVFG SSRTKSSASA SSHGRVRSED DQTSSNSNSG SPAASLGKTS SRIFGRADRG | 60 |
| ASPSPSTPHS PLPPTTPSKS TTNATDKPSQ PIPSAPSAPS AKTENQSAPA PSRQTPTPAT | 120 |
| DPATVSASPK PQVESARMPS SNLAPITTPV AVGATASKDS KADKSADNTP TSDNAAAGNV | 180 |
| KMPDPVKPLS INIPINQEVA AQPSAELPKK QPKSEVKSIN GSIKERDDLT PVGVSNTSSF | 240 |
| HSRTNADQYR ADYHPDNASK YANPNHYSAR PETYHRKSFS AGAPSLGEQT PAQSRRQSTS | 300 |
| EHDIGHTRGP TANLSNAVEA RASSQHWMSA EDQRLQEDAD KRRHWKRWGP YLSERQWGTV | 360 |
| REDYSSNGDA WSHFPHELAR SRTYRWGEDG LAGLSDNHCR MGLSLALWNG KDRMLKERLF | 420 |
| GLANNEGNHG EDVKELYWYL DSTPTHSYMK MLYKYPQESY PYELFVRESR NRSRDVAEFE | 480 |
| ITDTDLFDED KYWDVFVEYA KDKDEENAIS IRISAYNRGN EPADLHIIPQ LVFRNYWFWP | 540 |
| KEEPPKPKMK QSGDYVIQAD HPDLGRYHLY CSTSPAPSAP PAKRGQAPEP ISDEEVVPDL | 600 |
| LFTENETNFE RLYSGNNRNI YAKDAFHDHI VPCHRQEKDR PKAVEKTRKI KKTVVREERR | 660 |
| PVQRDPVAVP PKGNGASAEA EPEAAAAEKQ FGTVGPTQEQ EYEIVEIEEE VEEEETYFED | 720 |
| PDESKPPRDY VNPNKEGTKA GAHYVFRQVP PFGGCAVVRM KLTPKTPSQD PAIDDDEQFD | 780 |
| TTVEARRSEA DEFYAKLISG PMSDDMKNVM RQALAGMLWN KQFYMFIQPE WLRGDPGQPA | 840 |
| PPPERKRIRN HDWRHLHMED ILSMPDKWEY PFSAVWDSAF HCIPLAMVDP AFAKKQLDLF | 900 |
| TREWYMKPDG ALPAYEWNFS DVNPPVHAWA TFRVFKIERK MYGREDLDFL ERVFQKLLIN | 960 |
| FTWWVNRKDS SGSNVFEGGF LGLDNIGPFN RSEPLPTGGT LRQADGTAWM AFYALNMLNM | 1020 |
| ALELAKHNPT YEDIASKFFE HFIFISDAMS FHDTFDDEST SLWSDKDGFY YDAIQWGPGN | 1080 |
| SQVIPVKSLV GLIPLYATLT IEPSALKRFP SFAKRMQWFL DNRPEMADRN IANMKVGGRG | 1140 |
| DRRLLSMVSR ERLELILKRM LDENEFLSPH GVRSLSKDHQ KNPFHVNVGG EDFGVGYWPG | 1200 |
| DSHSPMFGGN SNWRGPIWIC VNFLLIESLQ RFYQYYGDSF KVECPTGSGD YMHLGHVAEE | 1260 |
| LSHRLLGIFL RDDQGRRANN GGIPLLDYAP EFRDLVHFYE FFHGDTGKGL GASHQCGWTG | 1320 |
| LIAYTIYSTG ASYGLAQTPR TPKSTAAHYF DEHLTEDGRS EAGSVPYSTA YSRPPSPDEL | 1380 |
| 1380 |
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH63 | 869 | 1322 | 2.4e-16 | 0.3649122807017544 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 282904 | Herpes_BLLF1 | 3.98e-10 | 12 | 324 | 406 | 742 | Herpes virus major outer envelope glycoprotein (BLLF1). This family consists of the BLLF1 viral late glycoprotein, also termed gp350/220. It is the most abundantly expressed glycoprotein in the viral envelope of the Herpesviruses and is the major antigen responsible for stimulating the production of neutralising antibodies in vivo. |
| 397353 | Glyco_hydro_63 | 8.59e-09 | 947 | 1238 | 185 | 434 | Glycosyl hydrolase family 63 C-terminal domain. This is a family of eukaryotic enzymes belonging to glycosyl hydrolase family 63. They catalyze the specific cleavage of the non-reducing terminal glucose residue from Glc(3)Man(9)GlcNAc(2). Mannosyl oligosaccharide glucosidase EC:3.2.1.106 is the first enzyme in the N-linked oligosaccharide processing pathway. This family represents the C-terminal catalytic domain. |
| 223039 | PHA03307 | 1.70e-08 | 10 | 337 | 90 | 402 | transcriptional regulator ICP4; Provisional |
| 282904 | Herpes_BLLF1 | 1.04e-07 | 10 | 306 | 484 | 770 | Herpes virus major outer envelope glycoprotein (BLLF1). This family consists of the BLLF1 viral late glycoprotein, also termed gp350/220. It is the most abundantly expressed glycoprotein in the viral envelope of the Herpesviruses and is the major antigen responsible for stimulating the production of neutralising antibodies in vivo. |
| 223021 | PHA03247 | 2.17e-06 | 75 | 321 | 2873 | 3100 | large tegument protein UL36; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CDI54668.1|GH0 | 0.0 | 1 | 1380 | 1 | 1419 |
| CDU25534.1|GH0 | 0.0 | 1 | 1380 | 1 | 1392 |
| CBQ68557.1|GH0 | 0.0 | 1 | 1380 | 1 | 1413 |
| SJX64191.1|GH0 | 0.0 | 1 | 1380 | 1 | 1418 |
| SAM84832.1|GH0 | 0.0 | 1 | 1380 | 1 | 1379 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| sp|Q04336|YM54_YEAST | 0.0 | 329 | 1341 | 12 | 953 | Uncharacterized protein YMR196W OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=YMR196W PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
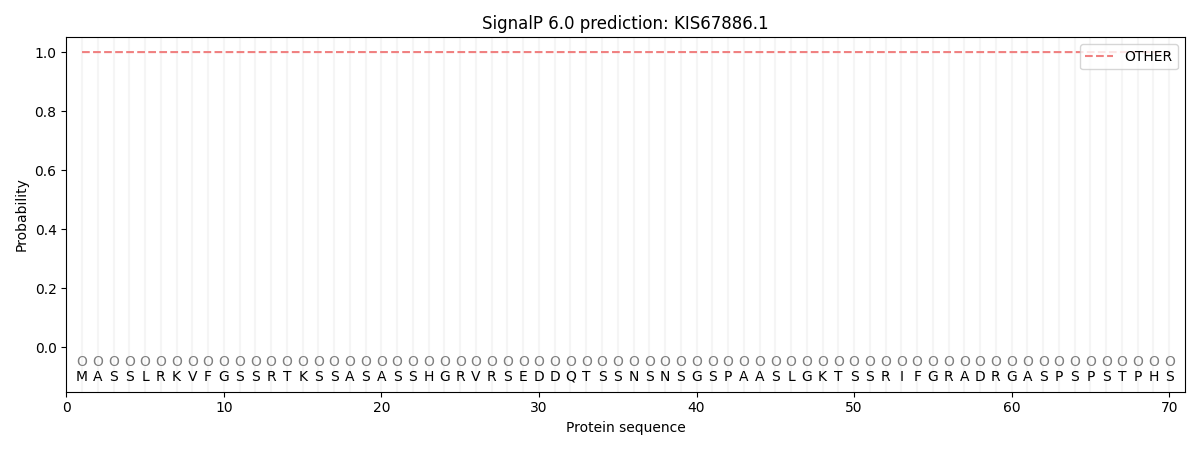
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000069 | 0.000000 |
