You are browsing environment: FUNGIDB
CAZyme Information: KID71695.1
You are here: Home > Sequence: KID71695.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Metarhizium anisopliae | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Clavicipitaceae; Metarhizium; Metarhizium anisopliae | |||||||||||
| CAZyme ID | KID71695.1 | |||||||||||
| CAZy Family | GT4 | |||||||||||
| CAZyme Description | glycoside hydrolase family 2 protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH2 | 17 | 545 | 4.6e-89 | 0.4867021276595745 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 236657 | PRK10150 | 2.86e-27 | 20 | 527 | 14 | 445 | beta-D-glucuronidase; Provisional |
| 225789 | LacZ | 3.56e-27 | 19 | 541 | 13 | 463 | Beta-galactosidase/beta-glucuronidase [Carbohydrate transport and metabolism]. |
| 236673 | ebgA | 4.00e-18 | 139 | 525 | 125 | 470 | cryptic beta-D-galactosidase subunit alpha; Reviewed |
| 397120 | Glyco_hydro_2_N | 3.91e-11 | 19 | 235 | 2 | 169 | Glycosyl hydrolases family 2, sugar binding domain. This family contains beta-galactosidase, beta-mannosidase and beta-glucuronidase activities and has a jelly-roll fold. The domain binds the sugar moiety during the sugar-hydrolysis reaction. |
| 397119 | Glyco_hydro_2_C | 3.35e-07 | 412 | 527 | 36 | 158 | Glycosyl hydrolases family 2, TIM barrel domain. This family contains beta-galactosidase, beta-mannosidase and beta-glucuronidase activities. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 692 | 1 | 692 | |
| 0.0 | 6 | 690 | 8 | 690 | |
| 0.0 | 4 | 692 | 2 | 690 | |
| 0.0 | 3 | 692 | 6 | 694 | |
| 0.0 | 5 | 689 | 2 | 684 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.20e-57 | 98 | 675 | 64 | 576 | Chain A, Glycosyl hydrolase family 2, sugar binding domain protein [Bacteroides cellulosilyticus DSM 14838],7SF2_B Chain B, Glycosyl hydrolase family 2, sugar binding domain protein [Bacteroides cellulosilyticus DSM 14838],7SF2_C Chain C, Glycosyl hydrolase family 2, sugar binding domain protein [Bacteroides cellulosilyticus DSM 14838],7SF2_D Chain D, Glycosyl hydrolase family 2, sugar binding domain protein [Bacteroides cellulosilyticus DSM 14838],7SF2_E Chain E, Glycosyl hydrolase family 2, sugar binding domain protein [Bacteroides cellulosilyticus DSM 14838],7SF2_F Chain F, Glycosyl hydrolase family 2, sugar binding domain protein [Bacteroides cellulosilyticus DSM 14838] |
|
| 1.07e-19 | 99 | 494 | 65 | 426 | Bacteroides uniforms beta-glucuronidase 2 bound to D-glucaro-1,5-lactone [Bacteroides uniformis str. 3978 T3 ii],6D50_B Bacteroides uniforms beta-glucuronidase 2 bound to D-glucaro-1,5-lactone [Bacteroides uniformis str. 3978 T3 ii] |
|
| 1.07e-19 | 99 | 494 | 57 | 418 | Crystal Structure of Bacteroides Uniformis beta-glucuronidase [Bacteroides uniformis str. 3978 T3 ii],5UJ6_B Crystal Structure of Bacteroides Uniformis beta-glucuronidase [Bacteroides uniformis str. 3978 T3 ii],6NZG_A Bacteroides uniformis beta-glucuronidase 2 covalently bound to cyclophellitol-6-carboxylate aziridine [Bacteroides uniformis],6NZG_B Bacteroides uniformis beta-glucuronidase 2 covalently bound to cyclophellitol-6-carboxylate aziridine [Bacteroides uniformis] |
|
| 3.25e-19 | 99 | 494 | 65 | 426 | D341A D367A calcium binding mutant of Bacteroides uniformis beta-glucuronidase 2 [Bacteroides uniformis str. 3978 T3 ii],6D8G_B D341A D367A calcium binding mutant of Bacteroides uniformis beta-glucuronidase 2 [Bacteroides uniformis str. 3978 T3 ii] |
|
| 2.05e-17 | 120 | 498 | 78 | 425 | Chain A, Glycosyl hydrolase family 2, sugar binding domain protein [Phocaeicola dorei],6ED1_B Chain B, Glycosyl hydrolase family 2, sugar binding domain protein [Phocaeicola dorei],6ED1_C Chain C, Glycosyl hydrolase family 2, sugar binding domain protein [Phocaeicola dorei],6ED1_D Chain D, Glycosyl hydrolase family 2, sugar binding domain protein [Phocaeicola dorei] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 9.83e-20 | 99 | 525 | 82 | 464 | Putative beta-glucuronidase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_21970 PE=2 SV=1 |
|
| 1.08e-18 | 120 | 538 | 57 | 429 | Beta-galactosidase OS=Thermoanaerobacter pseudethanolicus (strain ATCC 33223 / 39E) OX=340099 GN=lacZ PE=3 SV=2 |
|
| 5.61e-16 | 110 | 527 | 81 | 478 | Beta-glucuronidase OS=Felis catus OX=9685 GN=GUSB PE=1 SV=1 |
|
| 6.85e-15 | 110 | 527 | 81 | 478 | Beta-glucuronidase OS=Canis lupus familiaris OX=9615 GN=GUSB PE=1 SV=1 |
|
| 6.86e-15 | 121 | 527 | 100 | 479 | Beta-glucuronidase OS=Sus scrofa OX=9823 GN=GUSB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
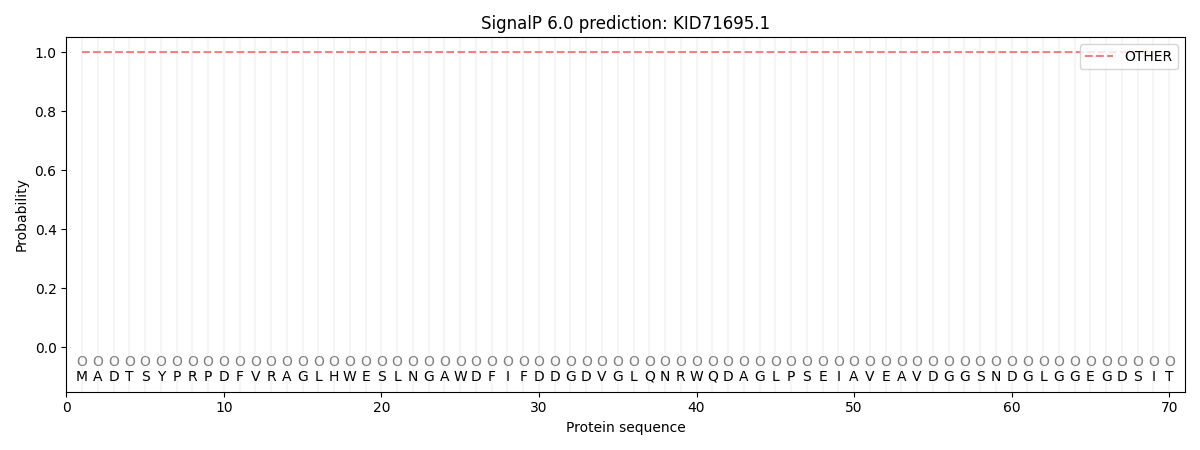
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000056 | 0.000001 |
