You are browsing environment: FUNGIDB
CAZyme Information: KFA50695.1
You are here: Home > Sequence: KFA50695.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Stachybotrys chartarum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Stachybotryaceae; Stachybotrys; Stachybotrys chartarum | |||||||||||
| CAZyme ID | KFA50695.1 | |||||||||||
| CAZy Family | PL3 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 2.4.1.173:23 | 2.4.1.-:3 |
|---|
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 340817 | GT1_Gtf-like | 2.01e-33 | 85 | 376 | 1 | 291 | UDP-glycosyltransferases and similar proteins. This family includes the Gtfs, a group of homologous glycosyltransferases involved in the final stages of the biosynthesis of antibiotics vancomycin and related chloroeremomycin. Gtfs transfer sugar moieties from an activated NDP-sugar donor to the oxidatively cross-linked heptapeptide core of vancomycin group antibiotics. The core structure is important for the bioactivity of the antibiotics. |
| 397255 | Glyco_transf_28 | 2.52e-16 | 87 | 220 | 1 | 138 | Glycosyltransferase family 28 N-terminal domain. The glycosyltransferase family 28 includes monogalactosyldiacylglycerol synthase (EC 2.4.1.46) and UDP-N-acetylglucosamine transferase (EC 2.4.1.-). This N-terminal domain contains the acceptor binding site and likely membrane association site. This family also contains a large number of proteins that probably have quite distinct activities. |
| 224732 | YjiC | 8.71e-12 | 87 | 375 | 4 | 285 | UDP:flavonoid glycosyltransferase YjiC, YdhE family [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 3.19e-197 | 12 | 584 | 31 | 796 | |
| 1.10e-190 | 12 | 585 | 31 | 847 | |
| 7.78e-186 | 12 | 584 | 31 | 849 | |
| 7.78e-186 | 12 | 584 | 31 | 849 | |
| 7.78e-186 | 12 | 584 | 31 | 849 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6.46e-28 | 92 | 357 | 27 | 297 | Sterol 3-beta-glucosyltransferase (ugt51) from Saccharomyces cerevisiae (strain ATCC 204508 / S288c) [Saccharomyces cerevisiae S288C],5XVM_B Sterol 3-beta-glucosyltransferase (ugt51) from Saccharomyces cerevisiae (strain ATCC 204508 / S288c) [Saccharomyces cerevisiae S288C] |
|
| 7.83e-28 | 92 | 357 | 47 | 317 | Sterol 3-beta-glucosyltransferase (ugt51) from Saccharomyces cerevisiae (strain ATCC 204508 / S288c): UDPG complex [Saccharomyces cerevisiae S288C],5GL5_B Sterol 3-beta-glucosyltransferase (ugt51) from Saccharomyces cerevisiae (strain ATCC 204508 / S288c): UDPG complex [Saccharomyces cerevisiae S288C] |
|
| 9.68e-06 | 85 | 334 | 1 | 248 | Crystal Structure of UDP-glucosyltransferase GtfB [Amycolatopsis orientalis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 8.51e-62 | 84 | 387 | 155 | 462 | Sterol 3-beta-glucosyltransferase UGT80B1 OS=Arabidopsis thaliana OX=3702 GN=UGT80B1 PE=1 SV=1 |
|
| 3.25e-59 | 66 | 340 | 168 | 461 | Sterol 3-beta-glucosyltransferase UGT80A2 OS=Arabidopsis thaliana OX=3702 GN=UGT80A2 PE=1 SV=1 |
|
| 2.78e-36 | 85 | 357 | 902 | 1177 | Sterol 3-beta-glucosyltransferase OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=atg26 PE=3 SV=2 |
|
| 1.60e-35 | 85 | 339 | 911 | 1173 | Sterol 3-beta-glucosyltransferase OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=atg26 PE=3 SV=1 |
|
| 3.77e-35 | 85 | 339 | 884 | 1146 | Sterol 3-beta-glucosyltransferase OS=Penicillium rubens (strain ATCC 28089 / DSM 1075 / NRRL 1951 / Wisconsin 54-1255) OX=500485 GN=atg26 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
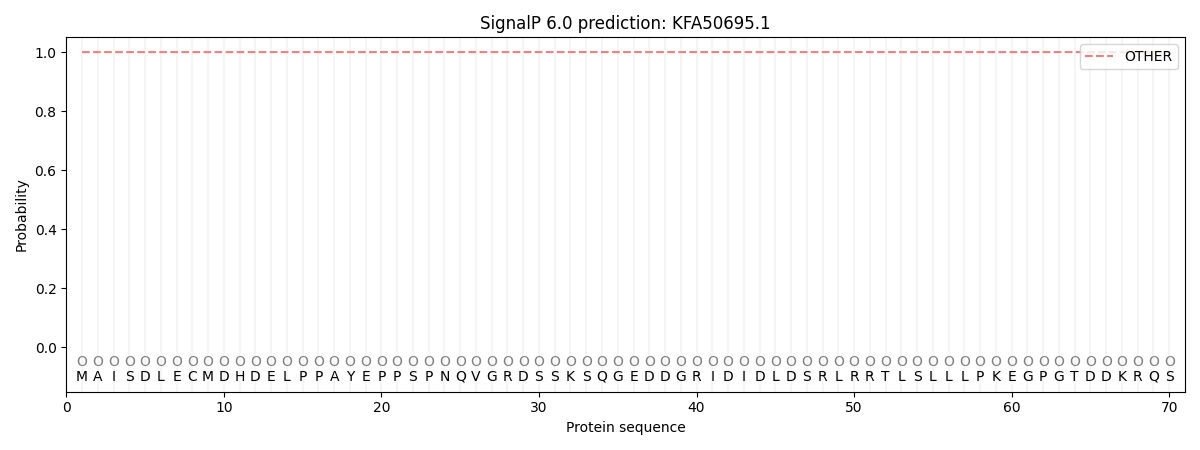
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000047 | 0.000000 |
