You are browsing environment: FUNGIDB
CAZyme Information: KFA50610.1
You are here: Home > Sequence: KFA50610.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Stachybotrys chartarum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Stachybotryaceae; Stachybotrys; Stachybotrys chartarum | |||||||||||
| CAZyme ID | KFA50610.1 | |||||||||||
| CAZy Family | GH128 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.8:67 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH11 | 37 | 215 | 1e-54 | 0.9830508474576272 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 395367 | Glyco_hydro_11 | 3.86e-74 | 35 | 213 | 1 | 175 | Glycosyl hydrolases family 11. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 6.42e-123 | 1 | 247 | 1 | 247 | |
| 7.47e-81 | 18 | 235 | 30 | 245 | |
| 4.06e-65 | 12 | 236 | 6 | 231 | |
| 4.61e-57 | 3 | 219 | 14 | 228 | |
| 1.23e-55 | 10 | 216 | 17 | 219 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.65e-52 | 35 | 217 | 15 | 191 | Chain A, Endo-1,4-beta-xylanase [Actinomycetia bacterium],7EO6_B Chain B, Endo-1,4-beta-xylanase [Actinomycetia bacterium] |
|
| 1.29e-51 | 35 | 216 | 12 | 189 | Crystal structure of family 11 xylanase in complex with inhibitor (XIP-I) [Talaromyces funiculosus] |
|
| 2.17e-51 | 28 | 216 | 12 | 206 | Xylanase 11C from Talaromyces cellulolyticus (formerly known as Acremonium cellulolyticus) [Talaromyces funiculosus],3WP3_B Xylanase 11C from Talaromyces cellulolyticus (formerly known as Acremonium cellulolyticus) [Talaromyces funiculosus] |
|
| 1.47e-50 | 35 | 216 | 11 | 189 | HIGH-RESOLUTION STRUCTURES OF XYLANASES FROM B. CIRCULANS AND T. HARZIANUM IDENTIFY A NEW FOLDING PATTERN AND IMPLICATIONS FOR THE ATOMIC BASIS OF THE CATALYSIS [Trichoderma harzianum] |
|
| 1.47e-50 | 35 | 216 | 12 | 189 | Chain A, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_B Chain B, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_C Chain C, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_D Chain D, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_E Chain E, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_F Chain F, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_G Chain G, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_H Chain H, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_I Chain I, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_J Chain J, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_K Chain K, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_L Chain L, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7.72e-55 | 5 | 216 | 5 | 222 | Effector Vd424Y OS=Verticillium dahliae (strain VdLs.17 / ATCC MYA-4575 / FGSC 10137) OX=498257 GN=VDAG_05042 PE=1 SV=1 |
|
| 8.25e-54 | 3 | 216 | 4 | 220 | Probable endo-1,4-beta-xylanase B OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=xlnB PE=3 SV=1 |
|
| 1.24e-53 | 1 | 216 | 3 | 222 | Endo-1,4-beta-xylanase C OS=Talaromyces funiculosus OX=28572 GN=xynC PE=1 SV=1 |
|
| 1.87e-52 | 3 | 216 | 4 | 220 | Probable endo-1,4-beta-xylanase B OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=xlnB PE=3 SV=1 |
|
| 1.87e-52 | 3 | 216 | 4 | 220 | Probable endo-1,4-beta-xylanase B OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=xlnB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
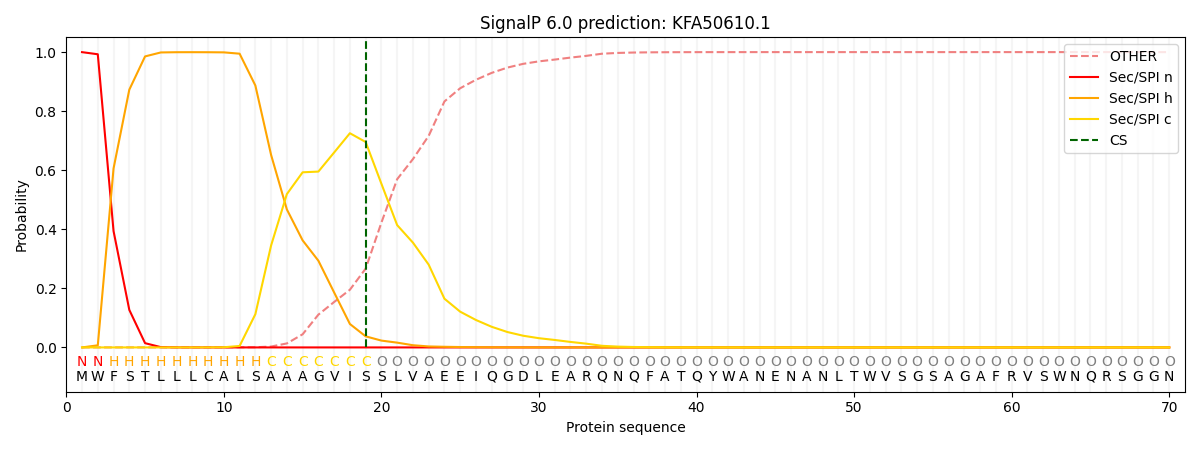
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000846 | 0.999142 | CS pos: 19-20. Pr: 0.6957 |
