You are browsing environment: FUNGIDB
CAZyme Information: KFA49202.1
You are here: Home > Sequence: KFA49202.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
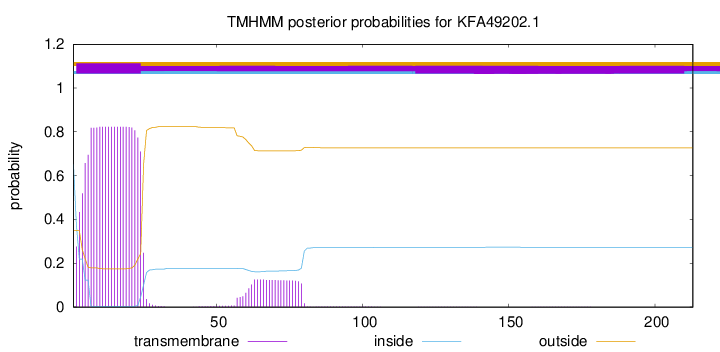
TMHMM annotations
Basic Information help
| Species | Stachybotrys chartarum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Stachybotryaceae; Stachybotrys; Stachybotrys chartarum | |||||||||||
| CAZyme ID | KFA49202.1 | |||||||||||
| CAZy Family | CE16 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 4.2.2.23:4 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL4 | 29 | 213 | 3.1e-53 | 0.3587174348697395 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 401282 | RhgB_N | 7.62e-79 | 22 | 213 | 1 | 213 | Rhamnogalacturonan lyase B, N-terminal. Members of this family are found in both fungi, bacteria and wood-eating arthropods. The domain is found at the N-terminus of rhamnogalacturonase B, a member of the polysaccharide lyase family 4. The domain adopts a structure consisting of a beta super-sandwich, with eighteen strands in two beta-sheets. The three domains of the whole protein rhamnogalacturonan lyase (RGL4), are involved in the degradation of rhamnogalacturonan-I, RG-I, an important pectic plant cell-wall polysaccharide. The active-site residues are a lysine at position 169 in UniProtKB:Q00019 and a histidine at 229, Lys169 is likely to be a proton abstractor, His229 a proton donor in the mechanism. The substrate is a disaccharide, and RGL4, in contrast to other rhamnogalacturonan hydrolases, cleaves the alpha-1,4 linkages of RG-I between Rha and GalUA through a beta-elimination resulting in a double bond in the nonreducing GalUA residue, and is thus classified as a polysaccharide lyase (PL). |
| 199907 | RGL4_N | 1.51e-34 | 23 | 213 | 47 | 236 | N-terminal catalytic domain of rhamnogalacturonan lyase, a family 4 polysaccharide lyase. The rhamnogalacturonan lyase of the polysaccharide lyase family 4 (RGL4) is involved in the degradation of RG (rhamnogalacturonan) type-I, an important pectic plant cell wall polysaccharide, by cleaving the alpha-1,4 glycoside bond between L-rhamnose and D-galacturonic acids in the backbone of RG type-I through a beta-elimination reaction. RGL4 consists of three domains, an N-terminal catalytic domain, a middle domain with a FNIII type fold and a C-terminal domain with a jelly roll fold; the middle and C-terminal domains are both putative carbohydrate binding modules. There are two types of RG lyases, which both cleave the alpha-1,4 bonds of the RG-I main chain (RG chain) through the beta-elimination reaction, but belong to two structurally unrelated polysaccharide lyase (PL) families, 4 and 11. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 3.80e-61 | 7 | 213 | 10 | 242 | |
| 2.59e-57 | 1 | 213 | 4 | 243 | |
| 8.35e-56 | 5 | 213 | 5 | 233 | |
| 8.35e-56 | 5 | 213 | 5 | 233 | |
| 8.34e-55 | 1 | 213 | 7 | 247 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.17e-50 | 36 | 213 | 41 | 213 | Rhamnogalacturonan lyase from Aspergillus aculeatus [Aspergillus aculeatus] |
|
| 1.63e-49 | 36 | 213 | 41 | 213 | Rhamnogalacturonan lyase from Aspergillus aculeatus K150A active site mutant [Aspergillus aculeatus],2XHN_B Rhamnogalacturonan lyase from Aspergillus aculeatus K150A active site mutant [Aspergillus aculeatus],3NJV_A Rhamnogalacturonan lyase from Aspergillus aculeatus K150A substrate complex [Aspergillus aculeatus] |
|
| 6.24e-49 | 36 | 213 | 41 | 213 | Rhamnogalacturonan Lyase from Aspergillus aculeatus mutant H210A [Aspergillus aculeatus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.60e-58 | 1 | 213 | 4 | 243 | Putative rhamnogalacturonase OS=Neurospora crassa (strain ATCC 24698 / 74-OR23-1A / CBS 708.71 / DSM 1257 / FGSC 987) OX=367110 GN=asd-1 PE=2 SV=1 |
|
| 1.48e-56 | 5 | 213 | 5 | 233 | Rhamnogalacturonate lyase A OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=rglA PE=2 SV=1 |
|
| 3.35e-54 | 7 | 213 | 7 | 233 | Probable rhamnogalacturonate lyase A OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=rglA PE=3 SV=1 |
|
| 6.57e-54 | 7 | 213 | 7 | 233 | Probable rhamnogalacturonate lyase A OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=rglA PE=3 SV=1 |
|
| 6.57e-54 | 7 | 213 | 7 | 233 | Probable rhamnogalacturonate lyase A OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=rglA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP

| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000222 | 0.999801 | CS pos: 20-21. Pr: 0.9804 |