You are browsing environment: FUNGIDB
CAZyme Information: KAG2020297.1
You are here: Home > Sequence: KAG2020297.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
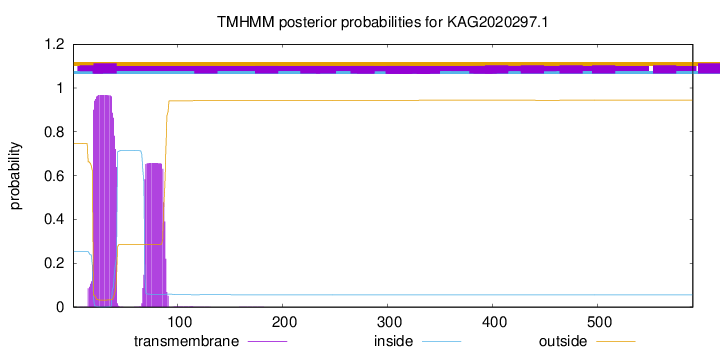
TMHMM annotations
Basic Information help
| Species | Coprinopsis cinerea | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Basidiomycota; Agaricomycetes; ; Psathyrellaceae; Coprinopsis; Coprinopsis cinerea | |||||||||||
| CAZyme ID | KAG2020297.1 | |||||||||||
| CAZy Family | GT15 | |||||||||||
| CAZyme Description | CEL4b mannanase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 73 | 510 | 3e-219 | 0.9977064220183486 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 226444 | COG3934 | 6.58e-15 | 69 | 307 | 4 | 187 | Endo-1,4-beta-mannosidase [Carbohydrate transport and metabolism]. |
| 200468 | AmyAc_maltase-like | 0.005 | 137 | 238 | 105 | 216 | Alpha amylase catalytic domain family found in maltase. Maltase (EC 3.2.1.20) hydrolyzes the terminal, non-reducing (1->4)-linked alpha-D-glucose residues in maltose, releasing alpha-D-glucose. The catalytic triad (DED) which is highly conserved in the other maltase group is not present in this subfamily. The Alpha-amylase family comprises the largest family of glycoside hydrolases (GH), with the majority of enzymes acting on starch, glycogen, and related oligo- and polysaccharides. These proteins catalyze the transformation of alpha-1,4 and alpha-1,6 glucosidic linkages with retention of the anomeric center. The protein is described as having 3 domains: A, B, C. A is a (beta/alpha) 8-barrel; B is a loop between the beta 3 strand and alpha 3 helix of A; C is the C-terminal extension characterized by a Greek key. The majority of the enzymes have an active site cleft found between domains A and B where a triad of catalytic residues (Asp, Glu and Asp) performs catalysis. Other members of this family have lost the catalytic activity as in the case of the human 4F2hc, or only have 2 residues that serve as the catalytic nucleophile and the acid/base, such as Thermus A4 beta-galactosidase with 2 Glu residues (GH42) and human alpha-galactosidase with 2 Asp residues (GH31). The family members are quite extensive and include: alpha amylase, maltosyltransferase, cyclodextrin glycotransferase, maltogenic amylase, neopullulanase, isoamylase, 1,4-alpha-D-glucan maltotetrahydrolase, 4-alpha-glucotransferase, oligo-1,6-glucosidase, amylosucrase, sucrose phosphorylase, and amylomaltase. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 23 | 591 | 1 | 498 | |
| 3.53e-211 | 23 | 588 | 27 | 595 | |
| 1.10e-175 | 70 | 587 | 1 | 539 | |
| 1.10e-175 | 70 | 587 | 1 | 539 | |
| 7.56e-136 | 27 | 540 | 9 | 543 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.01e-46 | 68 | 305 | 22 | 217 | Chain A, Gh5 Endo-beta-1,4-mannanase [Podospora anserina] |
|
| 2.12e-45 | 61 | 305 | 4 | 206 | Chain A, ENDO-1,4-B-D-MANNANASE [Trichoderma reesei],1QNP_A Chain A, Endo-1,4-b-d-mannanase [Trichoderma reesei],1QNQ_A Chain A, Endo-1,4-b-d-mannanase [Trichoderma reesei],1QNR_A Chain A, Endo-1,4-b-d-mannanase [Trichoderma reesei],1QNS_A Chain A, Endo-1,4-b-d-mannanase [Trichoderma reesei] |
|
| 5.76e-45 | 61 | 305 | 2 | 205 | The ligand-free structure of ManBK from Aspergillus niger BK01 [Aspergillus niger],3WH9_B The ligand-free structure of ManBK from Aspergillus niger BK01 [Aspergillus niger] |
|
| 7.42e-45 | 61 | 305 | 3 | 205 | Crtstal structure of glycoside hydrolase family 5 beta-mannanase from Talaromyces trachyspermus [Talaromyces trachyspermus] |
|
| 3.18e-39 | 61 | 465 | 4 | 298 | Crystal Structure Analysis of the Endo-1,4-beta-mannanase from Rhizomucor miehei [Rhizomucor miehei] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5.37e-48 | 61 | 316 | 97 | 313 | Mannan endo-1,4-beta-mannosidase F OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=manF PE=1 SV=2 |
|
| 5.37e-48 | 61 | 316 | 97 | 313 | Probable mannan endo-1,4-beta-mannosidase F OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=manF PE=3 SV=2 |
|
| 1.32e-46 | 61 | 316 | 94 | 310 | Mannan endo-1,4-beta-mannosidase F OS=Aspergillus clavatus (strain ATCC 1007 / CBS 513.65 / DSM 816 / NCTC 3887 / NRRL 1 / QM 1276 / 107) OX=344612 GN=manF PE=3 SV=1 |
|
| 1.65e-46 | 60 | 305 | 25 | 229 | Probable mannan endo-1,4-beta-mannosidase A OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=manA PE=3 SV=1 |
|
| 2.75e-46 | 61 | 316 | 114 | 330 | Probable mannan endo-1,4-beta-mannosidase F OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=manF PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
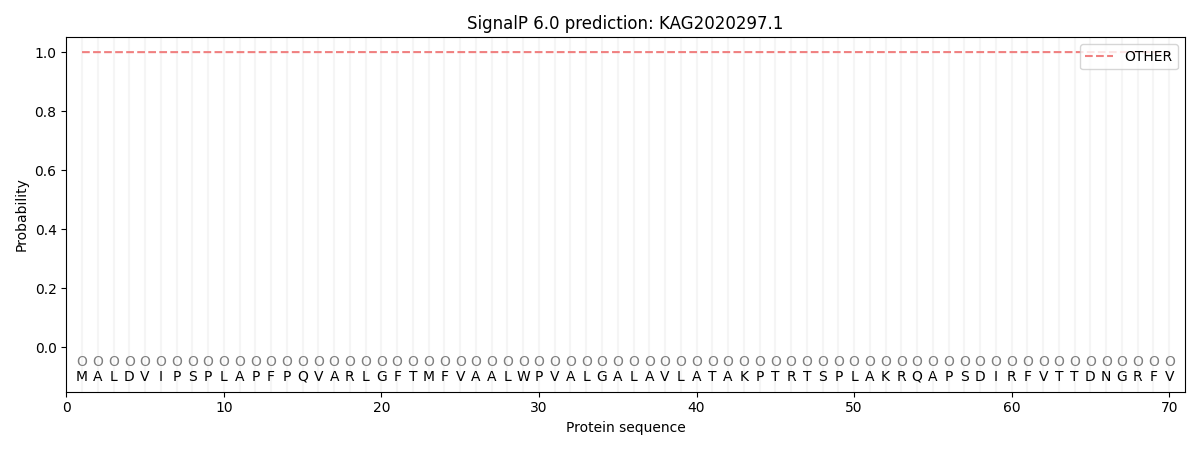
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.999945 | 0.000089 |