You are browsing environment: FUNGIDB
CAZyme Information: KAG2003658.1
You are here: Home > Sequence: KAG2003658.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
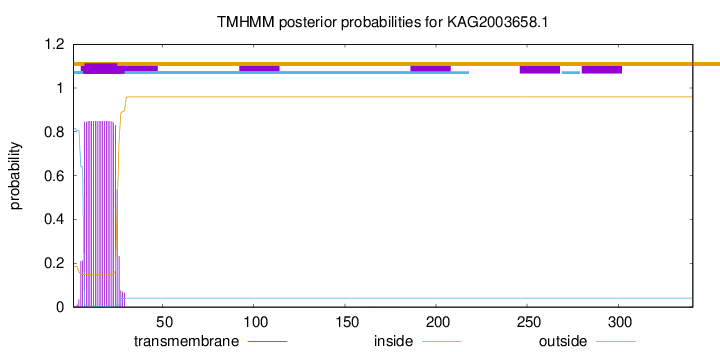
TMHMM annotations
Basic Information help
| Species | Coprinopsis cinerea | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Basidiomycota; Agaricomycetes; ; Psathyrellaceae; Coprinopsis; Coprinopsis cinerea | |||||||||||
| CAZyme ID | KAG2003658.1 | |||||||||||
| CAZy Family | AA3 | |||||||||||
| CAZyme Description | glycosyl hydrolase family 10 protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.8:31 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH10 | 29 | 332 | 4.6e-101 | 0.9735973597359736 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 395262 | Glyco_hydro_10 | 2.42e-151 | 22 | 332 | 1 | 310 | Glycosyl hydrolase family 10. |
| 214750 | Glyco_10 | 2.92e-121 | 67 | 330 | 1 | 263 | Glycosyl hydrolase family 10. |
| 226217 | XynA | 2.12e-92 | 4 | 332 | 7 | 339 | Endo-1,4-beta-xylanase, GH35 family [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.37e-161 | 26 | 336 | 36 | 346 | |
| 1.69e-134 | 21 | 333 | 18 | 330 | |
| 1.69e-134 | 21 | 333 | 18 | 330 | |
| 1.17e-133 | 22 | 333 | 86 | 397 | |
| 1.27e-131 | 26 | 333 | 84 | 391 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.34e-122 | 32 | 334 | 14 | 312 | The mutant crystal structure of b-1,4-Xylanase (XynAS9_V43P/G44E) from Streptomyces sp. 9 [Streptomyces sp.],3WUG_A The mutant crystal structure of b-1,4-Xylanase (XynAS9_V43P/G44E) with xylobiose from Streptomyces sp. 9 [Streptomyces sp.] |
|
| 3.11e-121 | 32 | 334 | 14 | 312 | The wild type crystal structure of b-1,4-Xylanase (XynAS9) from Streptomyces sp. 9 [Streptomyces sp.],3WUE_A The wild type crystal structure of b-1,4-Xylanase (XynAS9) with xylobiose from Streptomyces sp. 9 [Streptomyces sp.] |
|
| 4.38e-105 | 26 | 334 | 7 | 317 | Crystal structure of GH10 family xylanase XynAF1 from Aspergillus fumigatus Z5 [Aspergillus fumigatus Z5],6JDT_B Crystal structure of GH10 family xylanase XynAF1 from Aspergillus fumigatus Z5 [Aspergillus fumigatus Z5],6JDY_A Ligand complex structure of GH10 family xylanase XynAF1, soaking for 120 minutes [Aspergillus fumigatus Z5],6JDY_B Ligand complex structure of GH10 family xylanase XynAF1, soaking for 120 minutes [Aspergillus fumigatus Z5],6JDZ_A Ligand complex structure of GH10 family xylanase XynAF1, soaking for 20 minutes [Aspergillus fumigatus Z5],6JDZ_B Ligand complex structure of GH10 family xylanase XynAF1, soaking for 20 minutes [Aspergillus fumigatus Z5],6JE0_A Ligand complex structure of GH10 family xylanase XynAF1, soaking for 30 minutes [Aspergillus fumigatus Z5],6JE0_B Ligand complex structure of GH10 family xylanase XynAF1, soaking for 30 minutes [Aspergillus fumigatus Z5],6JE1_A Ligand complex structure of GH10 family xylanase XynAF1, soaking for 40 minutes [Aspergillus fumigatus Z5],6JE1_B Ligand complex structure of GH10 family xylanase XynAF1, soaking for 40 minutes [Aspergillus fumigatus Z5],6JE2_A Ligand complex structure of GH10 family xylanase XynAF1, soaking for 80 minutes [Aspergillus fumigatus Z5],6JE2_B Ligand complex structure of GH10 family xylanase XynAF1, soaking for 80 minutes [Aspergillus fumigatus Z5] |
|
| 2.64e-101 | 30 | 334 | 33 | 339 | GH10 endo-xylanase [Aspergillus aculeatus ATCC 16872],6Q8M_B GH10 endo-xylanase [Aspergillus aculeatus ATCC 16872],6Q8N_A GH10 endo-xylanase in complex with xylobiose epoxide inhibitor [Aspergillus aculeatus ATCC 16872],6Q8N_B GH10 endo-xylanase in complex with xylobiose epoxide inhibitor [Aspergillus aculeatus ATCC 16872] |
|
| 9.73e-100 | 34 | 332 | 15 | 310 | Beta-1,4-Glycanase Cex-Cd [Cellulomonas fimi],1FH7_A Crystal Structure Of The Xylanase Cex With Xylobiose- Derived Inhibitor Deoxynojirimycin [Cellulomonas fimi],1FH8_A Crystal Structure Of The Xylanase Cex With Xylobiose-derived Isofagomine Inhibitor [Cellulomonas fimi],1FH9_A Crystal Structure Of The Xylanase Cex With Xylobiose-derived Lactam Oxime Inhibitor [Cellulomonas fimi],1FHD_A Crystal Structure Of The Xylanase Cex With Xylobiose-derived Imidazole Inhibitor [Cellulomonas fimi],1J01_A Crystal Structure Of The Xylanase Cex With Xylobiose-Derived Inhibitor Isofagomine lactam [Cellulomonas fimi],2EXO_A Crystal Structure Of The Catalytic Domain Of The Beta-1,4- Glycanase Cex From Cellulomonas Fimi [Cellulomonas fimi],2XYL_A Cellulomonas Fimi XylanaseCELLULASE COMPLEXED WITH 2-Deoxy- 2-Fluoro-Xylobiose [Cellulomonas fimi] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.01e-135 | 21 | 333 | 18 | 330 | Endo-1,4-beta-xylanase OS=Agaricus bisporus OX=5341 GN=xlnA PE=2 SV=1 |
|
| 3.31e-120 | 25 | 333 | 97 | 405 | Endo-1,4-beta-xylanase A OS=Phanerodontia chrysosporium OX=2822231 GN=xynA PE=1 SV=1 |
|
| 2.25e-118 | 32 | 334 | 52 | 350 | Endo-1,4-beta-xylanase A OS=Streptomyces sp. OX=1931 GN=xynAS9 PE=1 SV=1 |
|
| 2.29e-118 | 25 | 333 | 87 | 396 | Endo-1,4-beta-xylanase C OS=Phanerodontia chrysosporium OX=2822231 GN=xynC PE=1 SV=1 |
|
| 4.93e-101 | 21 | 334 | 20 | 334 | Endo-1,4-beta-xylanase D OS=Talaromyces funiculosus OX=28572 GN=xynD PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
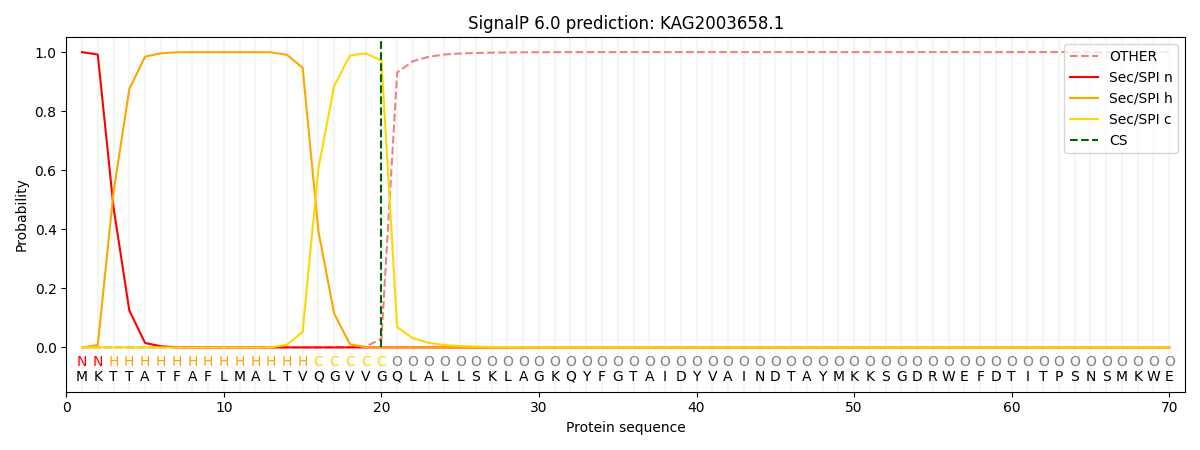
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.002164 | 0.997805 | CS pos: 20-21. Pr: 0.9713 |