You are browsing environment: FUNGIDB
CAZyme Information: KAG2003281.1
You are here: Home > Sequence: KAG2003281.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Coprinopsis cinerea | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Basidiomycota; Agaricomycetes; ; Psathyrellaceae; Coprinopsis; Coprinopsis cinerea | |||||||||||
| CAZyme ID | KAG2003281.1 | |||||||||||
| CAZy Family | AA2 | |||||||||||
| CAZyme Description | beta-glucosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.21:13 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH1 | 5 | 480 | 2.4e-165 | 0.986013986013986 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 274539 | BGL | 1.86e-170 | 7 | 477 | 1 | 426 | beta-galactosidase. |
| 395176 | Glyco_hydro_1 | 1.23e-164 | 6 | 479 | 5 | 446 | Glycosyl hydrolase family 1. |
| 225343 | BglB | 5.24e-146 | 4 | 479 | 2 | 448 | Beta-glucosidase/6-phospho-beta-glucosidase/beta-galactosidase [Carbohydrate transport and metabolism]. |
| 215435 | PLN02814 | 2.27e-119 | 6 | 477 | 28 | 476 | beta-glucosidase |
| 215455 | PLN02849 | 4.79e-117 | 7 | 477 | 31 | 476 | beta-glucosidase |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 2.06e-218 | 5 | 480 | 57 | 521 | |
| 2.82e-218 | 5 | 480 | 56 | 520 | |
| 1.53e-215 | 5 | 480 | 56 | 520 | |
| 1.08e-208 | 3 | 480 | 4 | 470 | |
| 9.07e-207 | 5 | 480 | 57 | 508 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.65e-206 | 1 | 480 | 4 | 462 | Chain A, Beta-glucosidase [Phanerodontia chrysosporium],2E3Z_B Chain B, Beta-glucosidase [Phanerodontia chrysosporium],2E40_A Chain A, Beta-glucosidase [Phanerodontia chrysosporium],2E40_B Chain B, Beta-glucosidase [Phanerodontia chrysosporium] |
|
| 6.18e-163 | 1 | 473 | 21 | 488 | Chain A, Beta-glucosidase [[Humicola] grisea var. thermoidea],4MDP_A Chain A, Beta-glucosidase [[Humicola] grisea var. thermoidea] |
|
| 1.41e-155 | 3 | 473 | 5 | 471 | Trichoderma harzianum GH1 beta-glucosidase ThBgl2 [Trichoderma harzianum] |
|
| 4.09e-154 | 3 | 473 | 18 | 484 | Chain A, Glycoside hydrolase family 1 [Trichoderma reesei QM6a],6KHT_B Chain B, Glycoside hydrolase family 1 [Trichoderma reesei QM6a],6KHT_C Chain C, Glycoside hydrolase family 1 [Trichoderma reesei QM6a] |
|
| 1.10e-147 | 6 | 479 | 3 | 461 | Crystal structure of the double mutant L167W / P172L of the beta-glucosidase from Trichoderma harzianum [Trichoderma harzianum],6EFU_B Crystal structure of the double mutant L167W / P172L of the beta-glucosidase from Trichoderma harzianum [Trichoderma harzianum] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7.62e-206 | 1 | 480 | 1 | 459 | Beta-glucosidase 1A OS=Phanerodontia chrysosporium OX=2822231 GN=BGL1A PE=1 SV=1 |
|
| 1.19e-204 | 5 | 482 | 10 | 476 | Beta-glucosidase 1B OS=Phanerodontia chrysosporium OX=2822231 GN=BGL1B PE=1 SV=1 |
|
| 5.80e-151 | 5 | 473 | 32 | 498 | Beta-glucosidase 24 OS=Oryza sativa subsp. japonica OX=39947 GN=BGLU24 PE=2 SV=1 |
|
| 1.58e-146 | 5 | 476 | 95 | 559 | Furostanol glycoside 26-O-beta-glucosidase OS=Hellenia speciosa OX=49577 GN=F26G PE=1 SV=1 |
|
| 2.85e-142 | 5 | 476 | 38 | 507 | Beta-glucosidase 12 OS=Oryza sativa subsp. japonica OX=39947 GN=BGLU12 PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
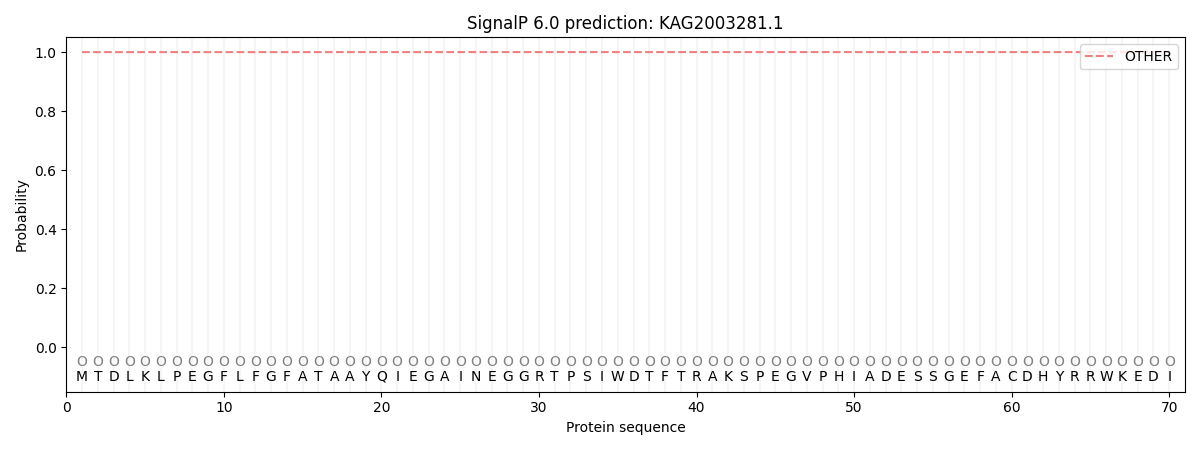
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000016 | 0.000026 |
