You are browsing environment: FUNGIDB
CAZyme Information: KAG2003144.1
You are here: Home > Sequence: KAG2003144.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Coprinopsis cinerea | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Basidiomycota; Agaricomycetes; ; Psathyrellaceae; Coprinopsis; Coprinopsis cinerea | |||||||||||
| CAZyme ID | KAG2003144.1 | |||||||||||
| CAZy Family | AA2 | |||||||||||
| CAZyme Description | cutinase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE5 | 31 | 198 | 9.4e-42 | 0.9841269841269841 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 395860 | Cutinase | 2.36e-51 | 31 | 199 | 2 | 173 | Cutinase. |
| 238382 | Lipase | 0.001 | 91 | 144 | 7 | 64 | Lipase. Lipases are esterases that can hydrolyze long-chain acyl-triglycerides into di- and monoglycerides, glycerol, and free fatty acids at a water/lipid interface. A typical feature of lipases is "interfacial activation", the process of becoming active at the lipid/water interface, although several examples of lipases have been identified that do not undergo interfacial activation . The active site of a lipase contains a catalytic triad consisting of Ser - His - Asp/Glu, but unlike most serine proteases, the active site is buried inside the structure. A "lid" or "flap" covers the active site, making it inaccessible to solvent and substrates. The lid opens during the process of interfacial activation, allowing the lipid substrate access to the active site. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 5.94e-72 | 1 | 199 | 1 | 200 | |
| 4.50e-51 | 7 | 199 | 5 | 197 | |
| 2.11e-50 | 14 | 195 | 16 | 196 | |
| 2.11e-50 | 14 | 195 | 16 | 196 | |
| 1.30e-48 | 31 | 196 | 42 | 207 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.10e-33 | 21 | 199 | 71 | 247 | Structure of cutinase from Trichoderma reesei in its native form. [Trichoderma reesei QM6a],4PSD_A Structure of Trichoderma reesei cutinase native form. [Trichoderma reesei QM6a],4PSE_A Trichoderma reesei cutinase in complex with a C11Y4 phosphonate inhibitor [Trichoderma reesei QM6a],4PSE_B Trichoderma reesei cutinase in complex with a C11Y4 phosphonate inhibitor [Trichoderma reesei QM6a] |
|
| 6.17e-26 | 20 | 192 | 6 | 199 | Chain A, CUTINASE [Fusarium vanettenii],1CUW_B Chain B, CUTINASE [Fusarium vanettenii] |
|
| 3.36e-25 | 19 | 199 | 1 | 198 | Chain A, cutinase [Malbranchea cinnamomea] |
|
| 1.25e-24 | 19 | 199 | 1 | 196 | Crystal structure of Aspergillus oryzae cutinase [Aspergillus oryzae] |
|
| 1.85e-24 | 20 | 192 | 6 | 199 | Chain A, CUTINASE [Fusarium vanettenii] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.07e-141 | 1 | 199 | 1 | 199 | Cutinase CUT1 OS=Coprinopsis cinerea OX=5346 GN=CUT1 PE=1 SV=1 |
|
| 3.74e-51 | 14 | 195 | 16 | 196 | Cutinase OS=Botryotinia fuckeliana OX=40559 GN=cutA PE=1 SV=1 |
|
| 2.36e-49 | 6 | 195 | 8 | 195 | Cutinase OS=Monilinia fructicola OX=38448 GN=CUT1 PE=2 SV=1 |
|
| 1.41e-37 | 1 | 198 | 1 | 200 | Cutinase pbc1 OS=Pyrenopeziza brassicae OX=76659 GN=pbc1 PE=1 SV=1 |
|
| 2.87e-35 | 11 | 195 | 58 | 243 | Cutinase cut1 OS=Trichoderma harzianum OX=5544 GN=cut1 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
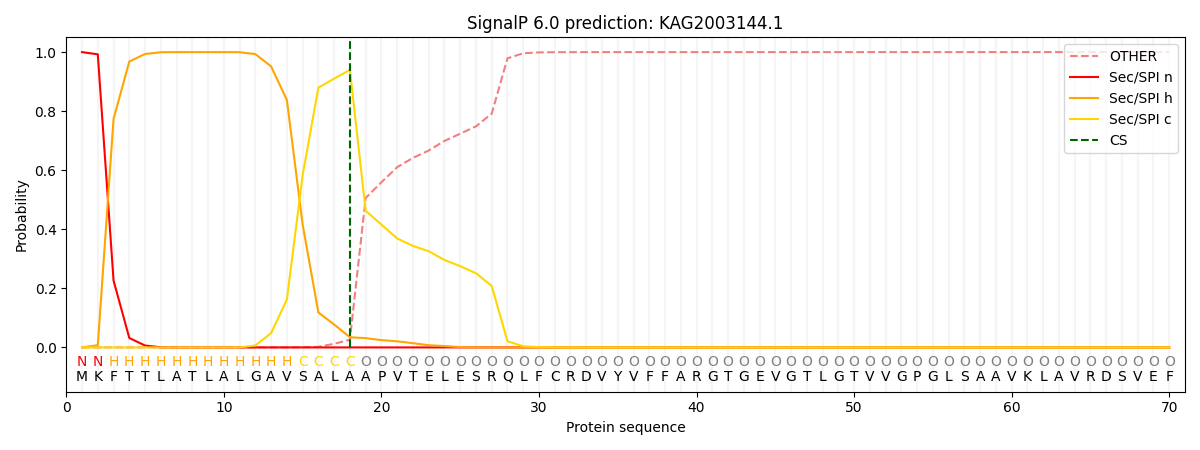
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000288 | 0.999683 | CS pos: 18-19. Pr: 0.9397 |
