You are browsing environment: FUNGIDB
CAZyme Information: KAG1702194.1
You are here: Home > Sequence: KAG1702194.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Phytophthora capsici | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Peronosporaceae; Phytophthora; Phytophthora capsici | |||||||||||
| CAZyme ID | KAG1702194.1 | |||||||||||
| CAZy Family | GH7 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH17 | 25 | 279 | 7.9e-22 | 0.977491961414791 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 227625 | Scw11 | 3.26e-22 | 23 | 275 | 45 | 304 | Exo-beta-1,3-glucanase, GH17 family [Carbohydrate transport and metabolism]. |
| 366033 | Glyco_hydro_17 | 1.03e-05 | 209 | 282 | 231 | 307 | Glycosyl hydrolases family 17. |
| 311610 | Glyco_hydro_53 | 3.97e-04 | 170 | 250 | 200 | 295 | Glycosyl hydrolase family 53. This domain belongs to family 53 of the glycosyl hydrolase classification. These enzymes are enzymes are endo-1,4- beta-galactanases (EC:3.2.1.89). The structure of this domain is known and has a TIM barrel fold. |
| 226385 | GanB | 8.37e-04 | 170 | 250 | 246 | 340 | Arabinogalactan endo-1,4-beta-galactosidase [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 2.11e-121 | 16 | 297 | 565 | 848 | |
| 4.32e-117 | 9 | 283 | 13 | 288 | |
| 4.46e-71 | 9 | 283 | 11 | 288 | |
| 1.12e-51 | 9 | 195 | 11 | 200 | |
| 8.04e-48 | 4 | 283 | 3 | 282 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.60e-11 | 25 | 285 | 39 | 297 | Crystal structure of glycoside hydrolase family 17 beta-1,3-glucanosyltransferase from Rhizomucor miehei [Rhizomucor miehei CAU432] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.28e-08 | 52 | 226 | 327 | 503 | Probable beta-glucosidase btgE OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=btgE PE=3 SV=1 |
|
| 4.90e-08 | 66 | 282 | 68 | 303 | Glucan 1,3-beta-glucosidase BGL2 OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=BGL2 PE=1 SV=2 |
|
| 5.18e-08 | 25 | 232 | 138 | 347 | Probable family 17 glucosidase SCW10 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=SCW10 PE=1 SV=1 |
|
| 1.60e-07 | 23 | 237 | 281 | 508 | Probable family 17 glucosidase SCW11 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=SCW11 PE=1 SV=1 |
|
| 2.09e-07 | 66 | 282 | 68 | 303 | Glucan 1,3-beta-glucosidase OS=Candida albicans OX=5476 GN=BGL2 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
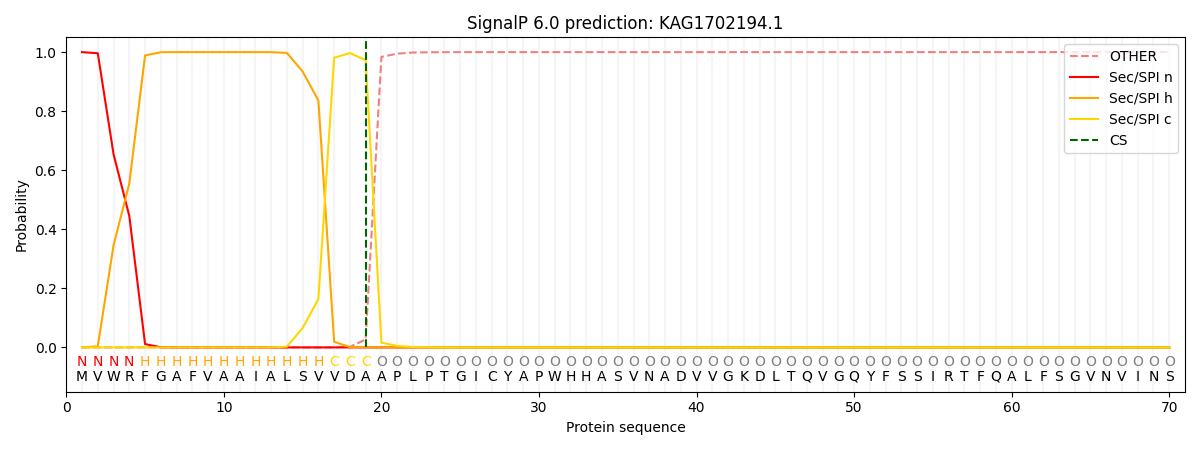
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000199 | 0.999754 | CS pos: 19-20. Pr: 0.9731 |
