You are browsing environment: FUNGIDB
CAZyme Information: KAG1698069.1
You are here: Home > Sequence: KAG1698069.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Phytophthora capsici | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Peronosporaceae; Phytophthora; Phytophthora capsici | |||||||||||
| CAZyme ID | KAG1698069.1 | |||||||||||
| CAZy Family | GH30 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL3 | 33 | 226 | 9.9e-80 | 0.979381443298969 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 397360 | Pectate_lyase | 1.61e-90 | 27 | 232 | 2 | 199 | Pectate lyase. |
| 411474 | fibronec_FbpA | 0.001 | 297 | 385 | 156 | 248 | LPXTG-anchored fibronectin-binding protein FbpA. FbpA, a fibronectin-binding protein described in Streptococcus pyogenes, has a YSIRK-type (crosswall-targeting) signal peptide and a C-terminal LPXTG motif for covalent attachment to the cell wall. It is unrelated to the PavA-like protein from Streptococcus gordonii (see BlastRule NBR009716) that was given the identical name, so the phase LPXTG-anchored is added to the protein name for clarity. |
| 411474 | fibronec_FbpA | 0.002 | 316 | 385 | 209 | 284 | LPXTG-anchored fibronectin-binding protein FbpA. FbpA, a fibronectin-binding protein described in Streptococcus pyogenes, has a YSIRK-type (crosswall-targeting) signal peptide and a C-terminal LPXTG motif for covalent attachment to the cell wall. It is unrelated to the PavA-like protein from Streptococcus gordonii (see BlastRule NBR009716) that was given the identical name, so the phase LPXTG-anchored is added to the protein name for clarity. |
| 237019 | PRK11907 | 0.010 | 314 | 386 | 34 | 107 | bifunctional 2',3'-cyclic-nucleotide 2'-phosphodiesterase/3'-nucleotidase. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 9.21e-125 | 1 | 265 | 1 | 262 | |
| 2.25e-116 | 21 | 253 | 28 | 258 | |
| 6.48e-107 | 1 | 165 | 1 | 165 | |
| 8.99e-106 | 21 | 253 | 28 | 258 | |
| 3.11e-97 | 17 | 261 | 16 | 281 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.44e-25 | 40 | 172 | 7 | 135 | Crystal Structure Of Pectate Lyase From Bacillus Sp. Strain Ksm-P15. [Bacillus sp. KSM-P15] |
|
| 4.00e-22 | 84 | 214 | 42 | 181 | Chain A, Endo-pectate lyase [Dickeya chrysanthemi],3B90_B Chain B, Endo-pectate lyase [Dickeya chrysanthemi] |
|
| 2.95e-21 | 84 | 214 | 160 | 299 | Chain A, Endo-pectate lyase [Dickeya chrysanthemi],3B4N_B Chain B, Endo-pectate lyase [Dickeya chrysanthemi],3B8Y_A Chain A, Endo-pectate lyase [Dickeya chrysanthemi],3B8Y_B Chain B, Endo-pectate lyase [Dickeya chrysanthemi] |
|
| 2.27e-19 | 34 | 186 | 6 | 156 | The liganded structure of C. bescii family 3 pectate lyase [Caldicellulosiruptor bescii DSM 6725],4EW9_B The liganded structure of C. bescii family 3 pectate lyase [Caldicellulosiruptor bescii DSM 6725] |
|
| 2.32e-19 | 34 | 186 | 7 | 157 | The crystal structure of family 3 pectate lyase from Caldicellulosiruptor bescii [Caldicellulosiruptor bescii],3T9G_B The crystal structure of family 3 pectate lyase from Caldicellulosiruptor bescii [Caldicellulosiruptor bescii] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7.50e-70 | 30 | 250 | 43 | 259 | Probable pectate lyase D OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=plyD PE=3 SV=1 |
|
| 3.48e-69 | 30 | 250 | 28 | 241 | Probable pectate lyase G OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=plyG PE=3 SV=1 |
|
| 5.77e-65 | 30 | 250 | 41 | 255 | Pectate lyase H OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=plyH PE=1 SV=1 |
|
| 9.27e-65 | 30 | 250 | 23 | 237 | Probable pectate lyase D OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=plyD PE=3 SV=1 |
|
| 9.27e-65 | 30 | 250 | 23 | 237 | Probable pectate lyase D OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=plyD PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
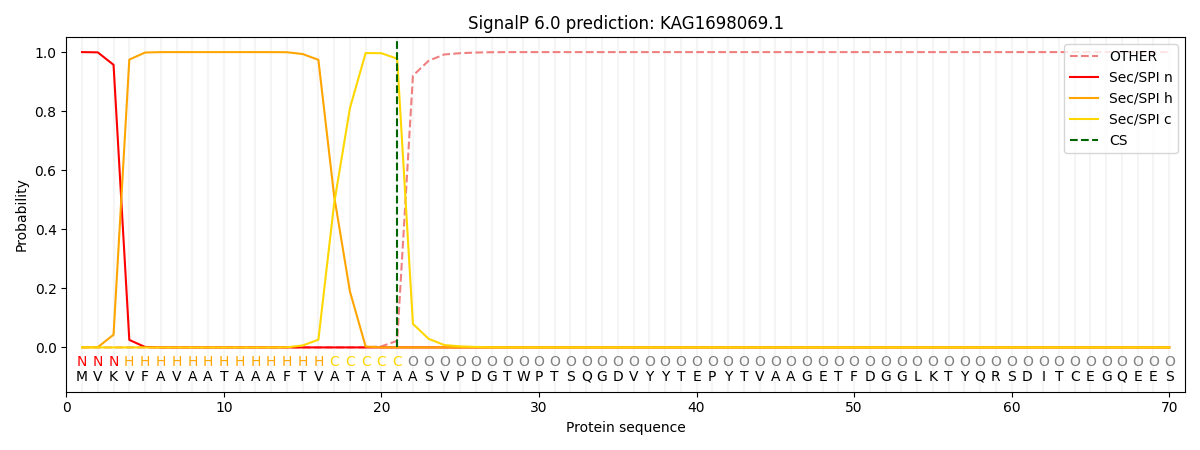
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000238 | 0.999756 | CS pos: 21-22. Pr: 0.9776 |
