You are browsing environment: FUNGIDB
CAZyme Information: KAF7618934.1
You are here: Home > Sequence: KAF7618934.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus flavus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus flavus | |||||||||||
| CAZyme ID | KAF7618934.1 | |||||||||||
| CAZy Family | CE12 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT76 | 26 | 454 | 1.2e-107 | 0.9877149877149877 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 282094 | Mannosyl_trans2 | 2.18e-33 | 78 | 454 | 57 | 439 | Mannosyltransferase (PIG-V). This is a family of eukaryotic ER membrane proteins that are involved in the synthesis of glycosylphosphatidylinositol (GPI), a glycolipid that anchors many proteins to the eukaryotic cell surface. Proteins in this family are involved in transferring the second mannose in the biosynthetic pathway of GPI. |
| 227829 | COG5542 | 6.07e-16 | 80 | 454 | 76 | 408 | Mannosyltransferase related to Gpi18 [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 454 | 1 | 452 | |
| 0.0 | 1 | 454 | 1 | 452 | |
| 0.0 | 1 | 454 | 1 | 452 | |
| 1.66e-309 | 1 | 454 | 1 | 434 | |
| 1.46e-234 | 118 | 454 | 14 | 350 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.94e-310 | 1 | 454 | 1 | 434 | GPI mannosyltransferase 2 OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=gpi18 PE=3 SV=1 |
|
| 3.45e-155 | 12 | 454 | 23 | 441 | GPI mannosyltransferase 2 OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=gpi18 PE=3 SV=1 |
|
| 2.55e-90 | 16 | 454 | 16 | 471 | GPI mannosyltransferase 2 OS=Chaetomium globosum (strain ATCC 6205 / CBS 148.51 / DSM 1962 / NBRC 6347 / NRRL 1970) OX=306901 GN=GPI18 PE=3 SV=1 |
|
| 1.04e-82 | 11 | 454 | 2 | 423 | GPI mannosyltransferase 2 OS=Gibberella zeae (strain ATCC MYA-4620 / CBS 123657 / FGSC 9075 / NRRL 31084 / PH-1) OX=229533 GN=GPI18 PE=3 SV=1 |
|
| 1.36e-44 | 21 | 454 | 6 | 357 | GPI mannosyltransferase 2 OS=Yarrowia lipolytica (strain CLIB 122 / E 150) OX=284591 GN=GPI18 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER

| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.993219 | 0.006794 |
TMHMM Annotations download full data without filtering help
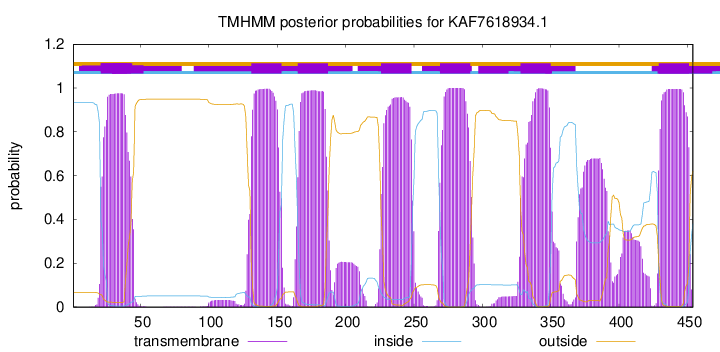
| Start | End |
|---|---|
| 21 | 43 |
| 131 | 153 |
| 165 | 187 |
| 226 | 248 |
| 269 | 291 |
| 328 | 350 |
| 429 | 451 |
