You are browsing environment: FUNGIDB
CAZyme Information: KAF6520365.1
You are here: Home > Sequence: KAF6520365.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Fusarium oxysporum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Nectriaceae; Fusarium; Fusarium oxysporum | |||||||||||
| CAZyme ID | KAF6520365.1 | |||||||||||
| CAZy Family | GH18 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 4.2.2.2:31 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL1 | 76 | 257 | 5e-85 | 0.994475138121547 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 214765 | Amb_all | 8.70e-71 | 77 | 260 | 2 | 189 | Amb_all domain. |
| 226384 | PelB | 6.76e-65 | 44 | 321 | 46 | 340 | Pectate lyase [Carbohydrate transport and metabolism]. |
| 366158 | Pec_lyase_C | 7.18e-45 | 93 | 257 | 39 | 211 | Pectate lyase. This enzyme forms a right handed beta helix structure. Pectate lyase is an enzyme involved in the maceration and soft rotting of plant tissue. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.10e-228 | 1 | 324 | 1 | 324 | |
| 1.75e-225 | 1 | 324 | 1 | 324 | |
| 5.01e-225 | 1 | 324 | 1 | 324 | |
| 5.85e-224 | 1 | 324 | 1 | 324 | |
| 1.18e-223 | 1 | 324 | 1 | 324 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 9.20e-38 | 38 | 259 | 6 | 248 | Crystal structure of pectate lyase Bsp165PelA from Bacillus sp. N165 [Bacillus sp. N16-5],3VMW_A Crystal structure of pectate lyase Bsp165PelA from Bacillus sp. N165 in complex with trigalacturonate [Bacillus sp. N16-5] |
|
| 1.86e-33 | 33 | 317 | 5 | 330 | Chain A, PECTATE LYASE E [Dickeya chrysanthemi] |
|
| 2.07e-24 | 94 | 302 | 92 | 303 | Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O88_A Chain A, Pectate Lyase C [Dickeya chrysanthemi],1O8D_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8E_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8F_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8G_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8H_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8I_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8J_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8K_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8L_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8M_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1PLU_A Chain A, Protein (pectate Lyase C) [Dickeya chrysanthemi],2PEC_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi] |
|
| 5.42e-24 | 94 | 302 | 92 | 303 | Chain A, Pectate lyase C [Dickeya chrysanthemi] |
|
| 6.02e-24 | 36 | 236 | 14 | 259 | Chain A, Pectate lyase A [Dickeya chrysanthemi],1OOC_B Chain B, Pectate lyase A [Dickeya chrysanthemi],1PE9_A Chain A, Pectate lyase A [Dickeya chrysanthemi],1PE9_B Chain B, Pectate lyase A [Dickeya chrysanthemi] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.66e-163 | 1 | 324 | 1 | 326 | Pectate lyase plyB OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=plyB PE=1 SV=1 |
|
| 2.10e-162 | 9 | 324 | 5 | 325 | Probable pectate lyase B OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=plyB PE=3 SV=1 |
|
| 9.76e-159 | 1 | 324 | 1 | 326 | Probable pectate lyase B OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=plyB PE=3 SV=1 |
|
| 9.76e-159 | 1 | 324 | 1 | 326 | Probable pectate lyase B OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=plyB PE=3 SV=1 |
|
| 1.29e-104 | 33 | 324 | 43 | 329 | Pectate lyase B OS=Colletotrichum gloeosporioides OX=474922 GN=PLB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
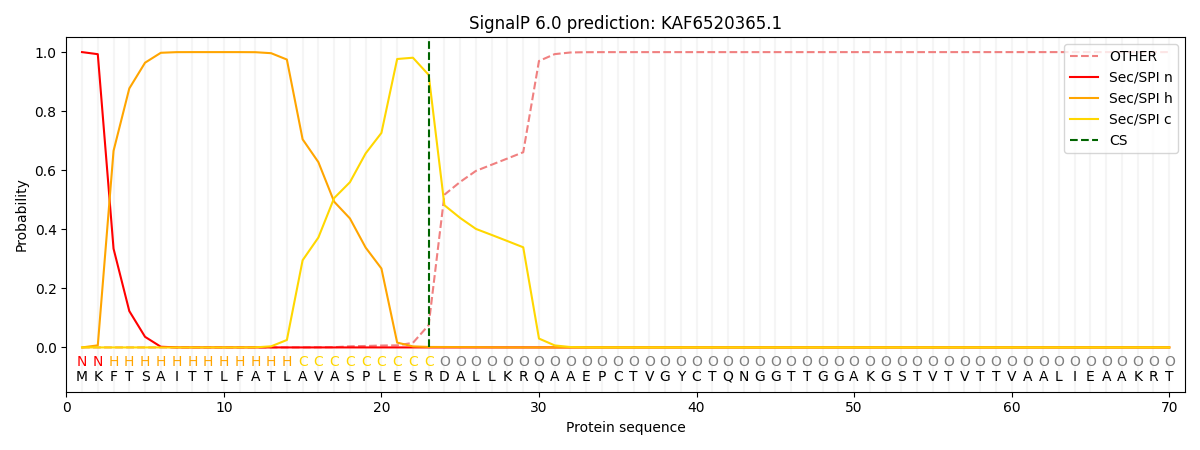
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000281 | 0.999681 | CS pos: 23-24. Pr: 0.9228 |
