You are browsing environment: FUNGIDB
CAZyme Information: KAF6517896.1
You are here: Home > Sequence: KAF6517896.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Fusarium oxysporum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Nectriaceae; Fusarium; Fusarium oxysporum | |||||||||||
| CAZyme ID | KAF6517896.1 | |||||||||||
| CAZy Family | GH10 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH88 | 134 | 490 | 8.7e-79 | 0.8723404255319149 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 271200 | LanM-like | 0.002 | 225 | 340 | 503 | 629 | Cyclases involved in the biosynthesis of class II lantibiotics, and similar proteins. LanM-like proteins. LanM is a bifunctional enzyme, involved in the synthesis of class II lantibiotics. It is responsible for both the dehydration and the cyclization of the precursor-peptide during lantibiotic synthesis. The C-terminal domain shows similarity to LanC, the cyclase component of the lan operon, but the N terminus seems to be unrelated to the dehydratase, LanB. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 501 | 1 | 537 | |
| 6.23e-286 | 1 | 500 | 1 | 531 | |
| 6.23e-286 | 1 | 500 | 1 | 531 | |
| 6.23e-286 | 1 | 500 | 1 | 531 | |
| 1.46e-284 | 1 | 500 | 1 | 531 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 8.26e-34 | 90 | 398 | 40 | 320 | Crystal structure of unsaturated glucuronyl hydrolase specific for heparin [Pedobacter heparinus DSM 2366] |
|
| 2.40e-23 | 99 | 339 | 44 | 282 | Crystal structure of unsaturated glucuronyl hydrolase from Streptcoccus agalactiae [Streptococcus agalactiae] |
|
| 2.43e-23 | 99 | 339 | 45 | 283 | Crystal structure of unsaturated glucuronyl hydrolase from Streptcoccus agalactiae [Streptococcus agalactiae serogroup III] |
|
| 1.10e-22 | 99 | 339 | 45 | 283 | Crystal structure of unsaturated glucuronyl hydrolase mutant D175N from Streptcoccus agalactiae [Streptococcus agalactiae serogroup III],3ANK_A Crystal structure of unsaturated glucuronyl hydrolase mutant D175N from Streptcoccus agalactiae complexed with dGlcA-GalNAc6S [Streptococcus agalactiae serogroup III] |
|
| 1.10e-22 | 99 | 339 | 45 | 283 | Crystal structure of unsaturated glucuronyl hydrolase mutant D115N from Streptcoccus agalactiae [Streptococcus agalactiae NEM316],3VXD_B Crystal structure of unsaturated glucuronyl hydrolase mutant D115N from Streptcoccus agalactiae [Streptococcus agalactiae NEM316],3VXD_C Crystal structure of unsaturated glucuronyl hydrolase mutant D115N from Streptcoccus agalactiae [Streptococcus agalactiae NEM316],3VXD_D Crystal structure of unsaturated glucuronyl hydrolase mutant D115N from Streptcoccus agalactiae [Streptococcus agalactiae NEM316] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.46e-32 | 123 | 490 | 77 | 389 | Unsaturated glucuronyl hydrolase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_21900 PE=1 SV=1 |
|
| 5.09e-29 | 99 | 418 | 46 | 337 | Unsaturated chondroitin disaccharide hydrolase OS=Streptococcus pyogenes serotype M1 OX=301447 GN=ugl PE=1 SV=1 |
|
| 1.09e-27 | 85 | 394 | 29 | 313 | Unsaturated chondroitin disaccharide hydrolase OS=Streptococcus pneumoniae (strain ATCC BAA-255 / R6) OX=171101 GN=ugl PE=1 SV=1 |
|
| 1.25e-22 | 99 | 339 | 45 | 283 | Unsaturated chondroitin disaccharide hydrolase OS=Streptococcus agalactiae serotype III (strain NEM316) OX=211110 GN=gbs1889 PE=1 SV=1 |
|
| 1.09e-21 | 101 | 417 | 20 | 310 | Unsaturated glucuronyl hydrolase OS=Bacillus sp. (strain GL1) OX=84635 GN=ugl PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
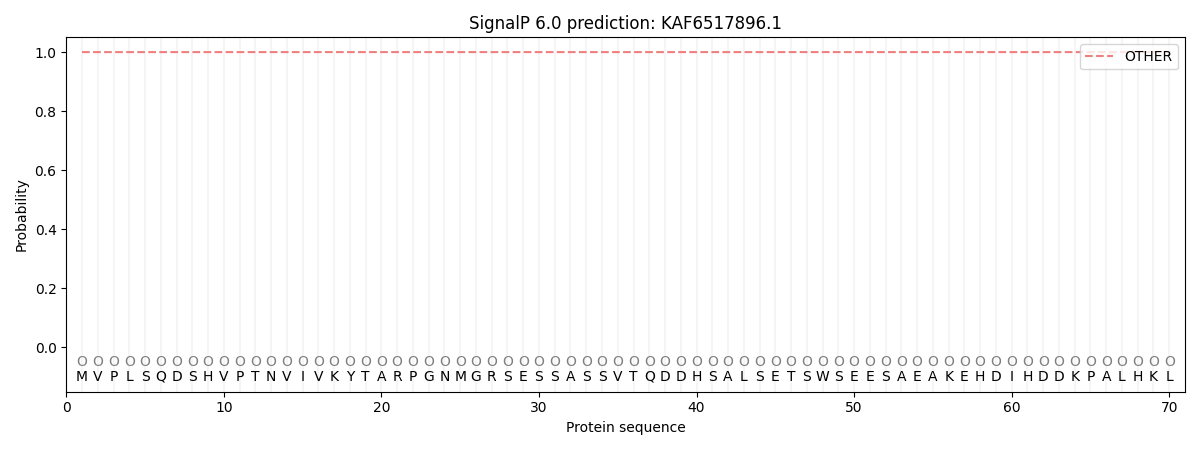
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000037 | 0.000001 |
