You are browsing environment: FUNGIDB
CAZyme Information: KAF5686023.1
You are here: Home > Sequence: KAF5686023.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Fusarium circinatum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Nectriaceae; Fusarium; Fusarium circinatum | |||||||||||
| CAZyme ID | KAF5686023.1 | |||||||||||
| CAZy Family | GH76 | |||||||||||
| CAZyme Description | isoamyl alcohol oxidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| AA7 | 118 | 564 | 3.3e-51 | 0.9432314410480349 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 225220 | COG2343 | 5.65e-41 | 691 | 820 | 1 | 128 | Uncharacterized conserved protein, DUF427 family [Function unknown]. |
| 398091 | NTP_transf_9 | 4.83e-38 | 720 | 812 | 1 | 93 | Domain of unknown function (DUF427). This domain contains a beta-tent fold. |
| 223354 | GlcD | 2.97e-16 | 118 | 353 | 30 | 266 | FAD/FMN-containing dehydrogenase [Energy production and conversion]. |
| 396238 | FAD_binding_4 | 5.70e-14 | 120 | 264 | 1 | 139 | FAD binding domain. This family consists of various enzymes that use FAD as a co-factor, most of the enzymes are similar to oxygen oxidoreductase. One of the enzymes Vanillyl-alcohol oxidase (VAO) has a solved structure, the alignment includes the FAD binding site, called the PP-loop, between residues 99-110. The FAD molecule is covalently bound in the known structure, however the residue that links to the FAD is not in the alignment. VAO catalyzes the oxidation of a wide variety of substrates, ranging form aromatic amines to 4-alkylphenols. Other members of this family include D-lactate dehydrogenase, this enzyme catalyzes the conversion of D-lactate to pyruvate using FAD as a co-factor; mitomycin radical oxidase, this enzyme oxidizes the reduced form of mitomycins and is involved in mitomycin resistance. This family includes MurB an UDP-N-acetylenolpyruvoylglucosamine reductase enzyme EC:1.1.1.158. This enzyme is involved in the biosynthesis of peptidoglycan. |
| 215242 | PLN02441 | 4.44e-05 | 120 | 297 | 65 | 242 | cytokinin dehydrogenase |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 2.41e-236 | 12 | 562 | 490 | 1034 | |
| 2.41e-236 | 12 | 562 | 490 | 1034 | |
| 9.98e-19 | 120 | 300 | 70 | 241 | |
| 8.53e-18 | 96 | 300 | 248 | 447 | |
| 8.53e-18 | 96 | 300 | 248 | 447 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.56e-66 | 1 | 571 | 6 | 552 | Crystal structure of VAO-type flavoprotein MtVAO615 at pH 7.5 from Myceliophthora thermophila C1 [Thermothelomyces thermophilus ATCC 42464],6F73_A Crystal structure of VAO-type flavoprotein MtVAO615 at pH 5.0 from Myceliophthora thermophila C1 [Thermothelomyces thermophilus ATCC 42464],6F73_B Crystal structure of VAO-type flavoprotein MtVAO615 at pH 5.0 from Myceliophthora thermophila C1 [Thermothelomyces thermophilus ATCC 42464] |
|
| 9.44e-55 | 20 | 562 | 28 | 567 | Crystal structure of VAO-type flavoprotein MtVAO713 from Myceliophthora thermophila C1 [Thermothelomyces thermophilus ATCC 42464],6F74_B Crystal structure of VAO-type flavoprotein MtVAO713 from Myceliophthora thermophila C1 [Thermothelomyces thermophilus ATCC 42464],6F74_C Crystal structure of VAO-type flavoprotein MtVAO713 from Myceliophthora thermophila C1 [Thermothelomyces thermophilus ATCC 42464],6F74_D Crystal structure of VAO-type flavoprotein MtVAO713 from Myceliophthora thermophila C1 [Thermothelomyces thermophilus ATCC 42464] |
|
| 6.43e-19 | 120 | 344 | 45 | 269 | The crystal structure of EncM H138T mutant [Streptomyces maritimus],6FYE_B The crystal structure of EncM H138T mutant [Streptomyces maritimus] |
|
| 1.26e-18 | 120 | 294 | 48 | 216 | Physcomitrella patens BBE-like 1 variant D396N [Physcomitrium patens],6EO5_B Physcomitrella patens BBE-like 1 variant D396N [Physcomitrium patens] |
|
| 1.26e-18 | 120 | 294 | 48 | 216 | Physcomitrella patens BBE-like 1 wild-type [Physcomitrium patens],6EO4_B Physcomitrella patens BBE-like 1 wild-type [Physcomitrium patens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.54e-67 | 11 | 562 | 14 | 554 | FAD-linked oxidoreductase apf9 OS=Gibberella fujikuroi (strain CBS 195.34 / IMI 58289 / NRRL A-6831) OX=1279085 GN=apf9 PE=1 SV=1 |
|
| 2.61e-67 | 7 | 562 | 12 | 532 | FAD-linked oxidoreductase ZEB1 OS=Gibberella zeae (strain ATCC MYA-4620 / CBS 123657 / FGSC 9075 / NRRL 31084 / PH-1) OX=229533 GN=ZEB1 PE=2 SV=2 |
|
| 8.03e-66 | 1 | 571 | 6 | 552 | VAO-type flavoprotein oxidase VAO615 OS=Myceliophthora thermophila (strain ATCC 42464 / BCRC 31852 / DSM 1799) OX=573729 GN=MYCTH_2305637 PE=1 SV=1 |
|
| 1.95e-64 | 3 | 562 | 7 | 560 | Uncharacterized FAD-linked oxidoreductase ARB_02478 OS=Arthroderma benhamiae (strain ATCC MYA-4681 / CBS 112371) OX=663331 GN=ARB_07056 PE=1 SV=2 |
|
| 8.81e-63 | 3 | 571 | 14 | 545 | FAD-linked oxidoreductase patO OS=Penicillium expansum OX=27334 GN=patO PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
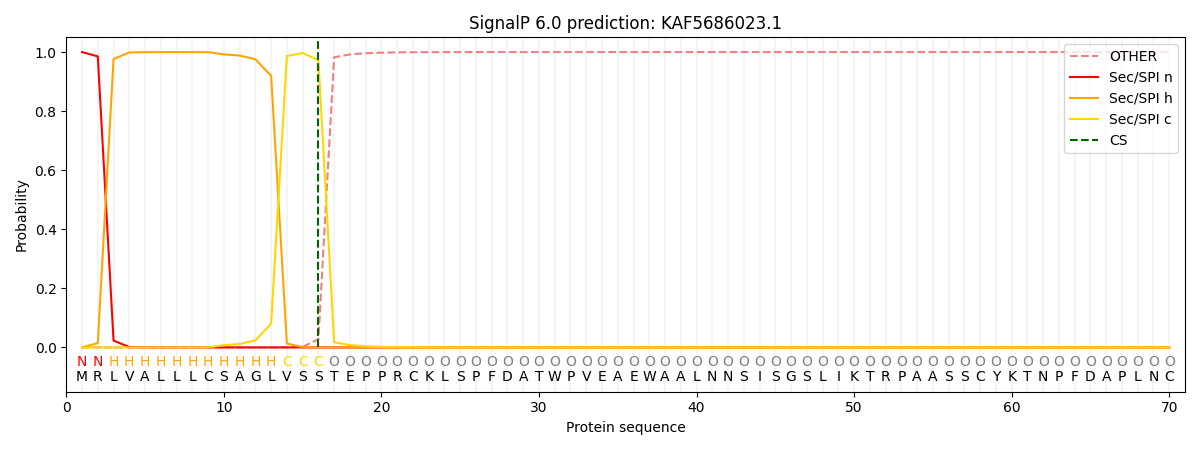
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000242 | 0.999748 | CS pos: 16-17. Pr: 0.9724 |
