You are browsing environment: FUNGIDB
CAZyme Information: KAE8541922.1
You are here: Home > Sequence: KAE8541922.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Cryptococcus cf. gattii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Arthropoda; Insecta; ; Eriococcidae; Cryptococcus; Cryptococcus cf. gattii | |||||||||||
| CAZyme ID | KAE8541922.1 | |||||||||||
| CAZy Family | GT2 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 974463; End:978192 Strand: - | |||||||||||
Full Sequence Download help
| MAPSQPSVRL LLLPFIKSRF LALLILPLVV LILYKNQLSK PPSLKVKVKE APQGGSWEAL | 60 |
| ADVDDDWGIH EGSWGGGVKD WFGWKDKGRR SIIITGGAGQ LAQALIPELI SNYTVHVQDI | 120 |
| VPRPPSLPSS VIYHRSSLSP AEAYKSIFPS NIFDGVIHLA GISLGAWCAA KEDQCHEVNV | 180 |
| GGTKALMDEV IAVNKQSKRG KMRKNVRVPW VILGSTMEVF GPEGTDEDSP KNPTSVIGRT | 240 |
| KLEAEQVFEE AILDGASEGI RGLVLRFPEV YGYSSASSIP EAFIPSLLTN SLTSLPIQYS | 300 |
| SDQIPYDLLH AEDAINGFVK AISYIEATPG GGVSSINLVS GQRTPARDIV ELVRKETTSM | 360 |
| SPVWDIGDNR NPSANEYSRA KAAEILSWQA EITPLVGLGK AITQLTEDIG TYSRSYLHEH | 420 |
| CPPSADFPAS DDDLAVSFIE DERNKPLWKL AGCTVNLGFN HEGWFRHLKC QDGKHCKVDD | 480 |
| VKVTALNWNQ STFIIEKVGK KQRERVVRVM LREQTGMGYL GMTKTEGEVG LELYKDSREG | 540 |
| QTVFDLEVRR DSSSLRLLVP DSEMQLHAVS NNTDGSTWFT VEPITRWVDP HFDMRINVLC | 600 |
| CPFEGDWPLL LDDYESADVR FGSTGQIPFD STRRSHLCAR AEQAVRYNFK HLSTARQAVN | 660 |
| KVTSTAGHAF FRETGPKTDP HPHGWALKDL PACWNDCGSP TVCVQTGNCK CVQADHCKPR | 720 |
| RENPLLKIYP SNPLHPDENK LTLGSLAGYS PVLAKAVEAI DWKDVLLPTA QAALEAHPDF | 780 |
| VKVHVADGYK GQEEIEAASC HKLQESDCFS ADSIMYRAFR HMSVPADEAE LVVVPAYQQC | 840 |
| EGTQFLLHDA MHHASETIQG VKNGEKKVAL VLTHDWGICV DFAWDIWSAR GERALHPDGI | 900 |
| LNNVLVWSVM GDYDSPCYRP HQDVVIPART CRSNTLRETF PNVGAIKPMR ERSNLLMWSG | 960 |
| TYSGTGKSER IRLTCNRGGA GDRELIKGGG KQSNFANSDY MKDLNNARFC PQPRGIAGWS | 1020 |
| SQTSDAIYAG CIPVFISEGT HYPFADFLDW SKLSVRVAPT ELDKIEKVLA AIPLSKVEEL | 1080 |
| QANLVSVREA FLYSGDQKPE DELERRGPMF FALHEAGMRI RTRYPVKGEE | 1130 |
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT47 | 781 | 1072 | 2e-42 | 0.9831081081081081 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 397245 | Exostosin | 1.63e-64 | 777 | 1072 | 1 | 290 | Exostosin family. The EXT family is a family of tumor suppressor genes. Mutations of EXT1 on 8q24.1, EXT2 on 11p11-13, and EXT3 on 19p have been associated with the autosomal dominant disorder known as hereditary multiple exostoses (HME). This is the most common known skeletal dysplasia. The chromosomal locations of other EXT genes suggest association with other forms of neoplasia. EXT1 and EXT2 have both been shown to encode a heparan sulphate polymerase with both D-glucuronyl (GlcA) and N-acetyl-D-glucosaminoglycan (GlcNAC) transferase activities. The nature of the defect in heparan sulphate biosynthesis in HME is unclear. |
| 223528 | WcaG | 2.84e-28 | 92 | 402 | 3 | 305 | Nucleoside-diphosphate-sugar epimerase [Cell wall/membrane/envelope biogenesis]. |
| 396097 | Epimerase | 6.97e-24 | 92 | 330 | 1 | 233 | NAD dependent epimerase/dehydratase family. This family of proteins utilize NAD as a cofactor. The proteins in this family use nucleotide-sugar substrates for a variety of chemical reactions. |
| 212494 | SDR_e | 3.30e-22 | 92 | 338 | 1 | 199 | extended (e) SDRs. Extended SDRs are distinct from classical SDRs. In addition to the Rossmann fold (alpha/beta folding pattern with a central beta-sheet) core region typical of all SDRs, extended SDRs have a less conserved C-terminal extension of approximately 100 amino acids. Extended SDRs are a diverse collection of proteins, and include isomerases, epimerases, oxidoreductases, and lyases; they typically have a TGXXGXXG cofactor binding motif. SDRs are a functionally diverse family of oxidoreductases that have a single domain with a structurally conserved Rossmann fold, an NAD(P)(H)-binding region, and a structurally diverse C-terminal region. Sequence identity between different SDR enzymes is typically in the 15-30% range; they catalyze a wide range of activities including the metabolism of steroids, cofactors, carbohydrates, lipids, aromatic compounds, and amino acids, and act in redox sensing. Classical SDRs have an TGXXX[AG]XG cofactor binding motif and a YXXXK active site motif, with the Tyr residue of the active site motif serving as a critical catalytic residue (Tyr-151, human 15-hydroxyprostaglandin dehydrogenase numbering). In addition to the Tyr and Lys, there is often an upstream Ser and/or an Asn, contributing to the active site; while substrate binding is in the C-terminal region, which determines specificity. The standard reaction mechanism is a 4-pro-S hydride transfer and proton relay involving the conserved Tyr and Lys, a water molecule stabilized by Asn, and nicotinamide. Atypical SDRs generally lack the catalytic residues characteristic of the SDRs, and their glycine-rich NAD(P)-binding motif is often different from the forms normally seen in classical or extended SDRs. Complex (multidomain) SDRs such as ketoreductase domains of fatty acid synthase have a GGXGXXG NAD(P)-binding motif and an altered active site motif (YXXXN). Fungal type ketoacyl reductases have a TGXXXGX(1-2)G NAD(P)-binding motif. |
| 187549 | Gne_like_SDR_e | 4.44e-17 | 91 | 321 | 2 | 224 | Escherichia coli Gne (a nucleoside-diphosphate-sugar 4-epimerase)-like, extended (e) SDRs. Nucleoside-diphosphate-sugar 4-epimerase has the characteristic active site tetrad and NAD-binding motif of the extended SDR, and is related to more specifically defined epimerases such as UDP-glucose 4 epimerase (aka UDP-galactose-4-epimerase), which catalyzes the NAD-dependent conversion of UDP-galactose to UDP-glucose, the final step in Leloir galactose synthesis. This subgroup includes Escherichia coli 055:H7 Gne, a UDP-GlcNAc 4-epimerase, essential for O55 antigen synthesis. Extended SDRs are distinct from classical SDRs. In addition to the Rossmann fold (alpha/beta folding pattern with a central beta-sheet) core region typical of all SDRs, extended SDRs have a less conserved C-terminal extension of approximately 100 amino acids. Extended SDRs are a diverse collection of proteins, and include isomerases, epimerases, oxidoreductases, and lyases; they typically have a TGXXGXXG cofactor binding motif. SDRs are a functionally diverse family of oxidoreductases that have a single domain with a structurally conserved Rossmann fold, an NAD(P)(H)-binding region, and a structurally diverse C-terminal region. Sequence identity between different SDR enzymes is typically in the 15-30% range; they catalyze a wide range of activities including the metabolism of steroids, cofactors, carbohydrates, lipids, aromatic compounds, and amino acids, and act in redox sensing. Classical SDRs have an TGXXX[AG]XG cofactor binding motif and a YXXXK active site motif, with the Tyr residue of the active site motif serving as a critical catalytic residue (Tyr-151, human 15-hydroxyprostaglandin dehydrogenase numbering). In addition to the Tyr and Lys, there is often an upstream Ser and/or an Asn, contributing to the active site; while substrate binding is in the C-terminal region, which determines specificity. The standard reaction mechanism is a 4-pro-S hydride transfer and proton relay involving the conserved Tyr and Lys, a water molecule stabilized by Asn, and nicotinamide. Atypical SDRs generally lack the catalytic residues characteristic of the SDRs, and their glycine-rich NAD(P)-binding motif is often different from the forms normally seen in classical or extended SDRs. Complex (multidomain) SDRs such as ketoreductase domains of fatty acid synthase have a GGXGXXG NAD(P)-binding motif and an altered active site motif (YXXXN). Fungal type ketoacyl reductases have a TGXXXGX(1-2)G NAD(P)-binding motif. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| KGB76296.2|GT47 | 0.0 | 1 | 1130 | 1 | 1130 |
| ADV23625.1|GT47 | 0.0 | 1 | 1130 | 1 | 1130 |
| AAW44475.1|GT47 | 0.0 | 1 | 1130 | 1 | 1132 |
| UOH83106.1|GT47 | 0.0 | 1 | 1130 | 1 | 1125 |
| AFR96705.1|GT47 | 0.0 | 1 | 1130 | 1 | 1125 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| sp|Q8S1X9|GT13_ORYSJ | 4.83e-14 | 827 | 1101 | 104 | 381 | Probable glucuronosyltransferase Os01g0926400 OS=Oryza sativa subsp. japonica OX=39947 GN=Os01g0926400 PE=2 SV=1 |
| sp|Q3EAR7|GLYT2_ARATH | 1.83e-12 | 999 | 1101 | 348 | 450 | Probable glycosyltransferase At3g42180 OS=Arabidopsis thaliana OX=3702 GN=At3g42180 PE=2 SV=2 |
| sp|Q94AA9|XGD1_ARATH | 1.89e-11 | 999 | 1101 | 378 | 480 | Xylogalacturonan beta-1,3-xylosyltransferase OS=Arabidopsis thaliana OX=3702 GN=XGD1 PE=1 SV=2 |
| sp|Q3E9A4|GLYT5_ARATH | 2.24e-11 | 997 | 1103 | 343 | 449 | Probable glycosyltransferase At5g20260 OS=Arabidopsis thaliana OX=3702 GN=At5g20260 PE=3 SV=3 |
| sp|Q6NMM8|F8H_ARATH | 3.97e-11 | 806 | 1101 | 133 | 438 | Probable glucuronoxylan glucuronosyltransferase F8H OS=Arabidopsis thaliana OX=3702 GN=F8H PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
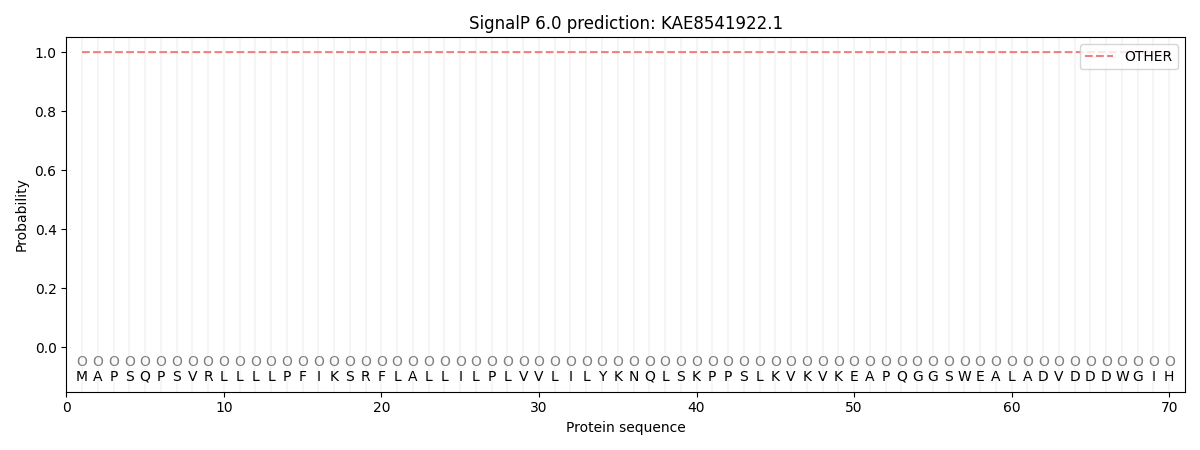
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.999959 | 0.000061 |

