You are browsing environment: FUNGIDB
CAZyme Information: KAE8541709.1
You are here: Home > Sequence: KAE8541709.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Cryptococcus cf. gattii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Arthropoda; Insecta; ; Eriococcidae; Cryptococcus; Cryptococcus cf. gattii | |||||||||||
| CAZyme ID | KAE8541709.1 | |||||||||||
| CAZy Family | GT15 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH47 | 223 | 760 | 5e-144 | 0.9910313901345291 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 396217 | Glyco_hydro_47 | 3.64e-177 | 223 | 760 | 2 | 451 | Glycosyl hydrolase family 47. Members of this family are alpha-mannosidases that catalyze the hydrolysis of the terminal 1,2-linked alpha-D-mannose residues in the oligo-mannose oligosaccharide Man(9)(GlcNAc)(2). |
| 240427 | PTZ00470 | 7.67e-118 | 159 | 765 | 21 | 521 | glycoside hydrolase family 47 protein; Provisional |
| 271199 | LanC_SerThrkinase | 0.005 | 332 | 433 | 94 | 188 | Lanthionine synthetase C-like domain associated with serine/threonine kinases. Some members of this subgroup lack the zinc binding site and the active site residues, and therefore are most likely inactive. The function of this domain is unknown. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 851 | 14 | 864 | |
| 0.0 | 1 | 851 | 1 | 851 | |
| 0.0 | 1 | 851 | 1 | 851 | |
| 0.0 | 1 | 851 | 14 | 865 | |
| 0.0 | 1 | 851 | 14 | 865 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.23e-72 | 223 | 760 | 13 | 449 | Crystal structure of the class I human endoplasmic reticulum 1,2-alpha-mannosidase and Man9GlcNAc2-PA complex [Homo sapiens] |
|
| 1.44e-72 | 223 | 760 | 18 | 454 | Crystal Structure Of Human Class I Alpha1,2-Mannosidase [Homo sapiens] |
|
| 1.52e-72 | 223 | 760 | 18 | 454 | Crystal Structure Of Human Class I Alpha1,2-Mannosidase In Complex With 1-Deoxymannojirimycin [Homo sapiens],1FO3_A Crystal Structure Of Human Class I Alpha1,2-Mannosidase In Complex With Kifunensine [Homo sapiens] |
|
| 6.85e-72 | 223 | 760 | 13 | 449 | Crystal structure of the class I human endoplasmic reticulum 1,2-alpha-mannosidase T688A mutant and Thio-disaccharide substrate analog complex [Homo sapiens],5KK7_B Crystal structure of the class I human endoplasmic reticulum 1,2-alpha-mannosidase T688A mutant and Thio-disaccharide substrate analog complex [Homo sapiens] |
|
| 1.10e-71 | 223 | 760 | 96 | 532 | Crystal Structure Of Human Class I alpha-1,2-Mannosidase In Complex With Thio-Disaccharide Substrate Analogue [Homo sapiens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.75e-69 | 223 | 760 | 257 | 693 | Endoplasmic reticulum mannosyl-oligosaccharide 1,2-alpha-mannosidase OS=Homo sapiens OX=9606 GN=MAN1B1 PE=1 SV=2 |
|
| 8.95e-67 | 214 | 760 | 206 | 651 | Endoplasmic reticulum mannosyl-oligosaccharide 1,2-alpha-mannosidase OS=Rattus norvegicus OX=10116 GN=Man1b1 PE=2 SV=2 |
|
| 1.17e-65 | 214 | 760 | 207 | 652 | Endoplasmic reticulum mannosyl-oligosaccharide 1,2-alpha-mannosidase OS=Mus musculus OX=10090 GN=Man1b1 PE=1 SV=1 |
|
| 1.92e-65 | 214 | 760 | 97 | 535 | Mannosyl-oligosaccharide 1,2-alpha-mannosidase MNS2 OS=Arabidopsis thaliana OX=3702 GN=MNS2 PE=1 SV=1 |
|
| 3.65e-64 | 211 | 772 | 93 | 546 | Mannosyl-oligosaccharide 1,2-alpha-mannosidase MNS1 OS=Arabidopsis thaliana OX=3702 GN=MNS1 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
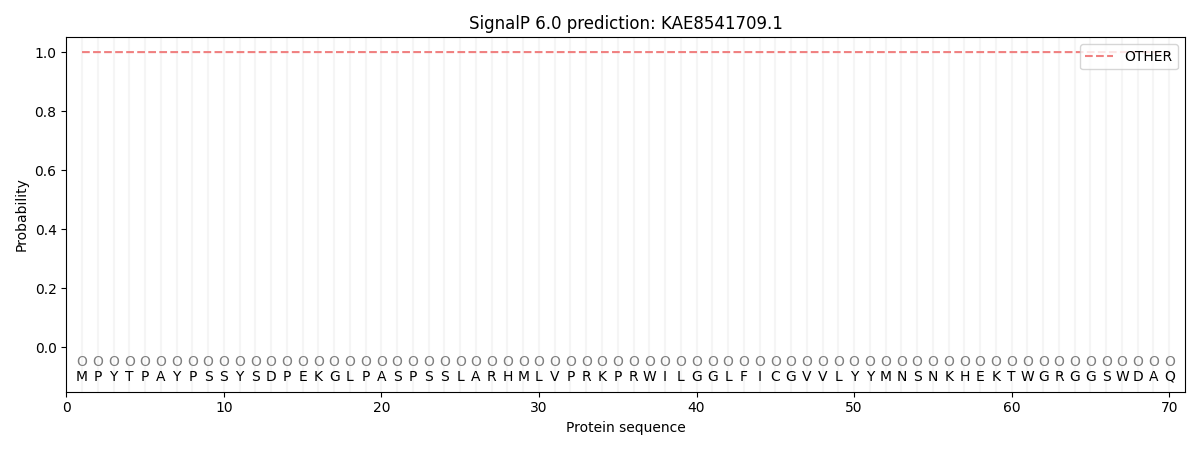
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000035 | 0.000012 |
