You are browsing environment: FUNGIDB
CAZyme Information: KAA8903558.1
You are here: Home > Sequence: KAA8903558.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Diutina rugosa | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Saccharomycetes; ; NA; Diutina; Diutina rugosa | |||||||||||
| CAZyme ID | KAA8903558.1 | |||||||||||
| CAZy Family | GT20 | |||||||||||
| CAZyme Description | Endo-1,3(4)-beta-glucanase [Source:UniProtKB/TrEMBL;Acc:A0A642UQK4] | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.39:20 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH81 | 634 | 1324 | 1.6e-219 | 0.9823151125401929 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 227785 | Acf2 | 0.0 | 616 | 1331 | 62 | 757 | Endoglucanase Acf2 [Carbohydrate transport and metabolism]. |
| 407548 | Glyco_hydro81C | 0.0 | 967 | 1323 | 1 | 349 | Glycosyl hydrolase family 81 C-terminal domain. Family of eukaryotic beta-1,3-glucanases. Within the Aspergillus fumigatus protein, two perfectly conserved Glu residues (E550 or E554) have been proposed as putative nucleophiles of the active site of the Engl1 endoglucanase, while the proton donor would be D475. The endo-beta-1,3-glucanase activity is essential for efficient spore release. This entry represents the helical C-terminal domain. |
| 397619 | Glyco_hydro_81 | 2.41e-101 | 634 | 960 | 5 | 318 | Glycosyl hydrolase family 81. Family of eukaryotic beta-1,3-glucanases. Within the Aspergillus fumigatus protein ENGL1, two perfectly conserved Glu residues (E550 or E554) have been proposed as putative nucleophiles of the active site of the Engl1 endoglucanase, while the proton donor would be D475. The endo-beta-1,3-glucanase activity is essential for efficient spore release. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 365 | 1330 | 168 | 1055 | |
| 6.48e-314 | 604 | 1332 | 434 | 1158 | |
| 4.20e-312 | 590 | 1333 | 319 | 1055 | |
| 2.07e-308 | 606 | 1333 | 425 | 1145 | |
| 3.26e-298 | 591 | 1329 | 356 | 1091 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.44e-94 | 608 | 1327 | 5 | 716 | The structure of a glycoside hydrolase family 81 endo-[beta]-1,3-glucanase [Rhizomucor miehei],4K3A_B The structure of a glycoside hydrolase family 81 endo-[beta]-1,3-glucanase [Rhizomucor miehei] |
|
| 2.51e-94 | 608 | 1327 | 30 | 741 | Crystal structure of GH family 81 beta-1,3-glucanase from Rhizomucr miehei complexed with laminaripentaose [Rhizomucor miehei],5XBZ_B Crystal structure of GH family 81 beta-1,3-glucanase from Rhizomucr miehei complexed with laminaripentaose [Rhizomucor miehei],5XC2_A Crystal structure of GH family 81 beta-1,3-glucanase from Rhizomucr miehei complexed with laminarihexaose [Rhizomucor miehei],5XC2_B Crystal structure of GH family 81 beta-1,3-glucanase from Rhizomucr miehei complexed with laminarihexaose [Rhizomucor miehei],5XC2_C Crystal structure of GH family 81 beta-1,3-glucanase from Rhizomucr miehei complexed with laminarihexaose [Rhizomucor miehei],5XC2_D Crystal structure of GH family 81 beta-1,3-glucanase from Rhizomucr miehei complexed with laminarihexaose [Rhizomucor miehei] |
|
| 1.46e-91 | 608 | 1327 | 5 | 716 | The structure of a glycoside hydrolase family 81 endo-[beta]-1,3-glucanase [Rhizomucor miehei],4K35_B The structure of a glycoside hydrolase family 81 endo-[beta]-1,3-glucanase [Rhizomucor miehei] |
|
| 2.44e-27 | 887 | 1306 | 242 | 637 | Chain A, Glycoside hydrolase family 81 [Acetivibrio thermocellus ATCC 27405] |
|
| 1.87e-20 | 844 | 1301 | 219 | 656 | Chain A, Glycoside Hydrolase [Halalkalibacterium halodurans C-125] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.86e-296 | 604 | 1332 | 422 | 1144 | Endo-1,3(4)-beta-glucanase 1 OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=ENG1 PE=1 SV=1 |
|
| 3.10e-240 | 614 | 1331 | 405 | 1114 | Endo-1,3(4)-beta-glucanase 1 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=DSE4 PE=1 SV=1 |
|
| 2.26e-165 | 609 | 1329 | 46 | 737 | Primary septum endo-1,3(4)-beta-glucanase OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=eng1 PE=3 SV=1 |
|
| 1.42e-161 | 593 | 1330 | 58 | 779 | Endo-1,3(4)-beta-glucanase 2 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=ACF2 PE=1 SV=1 |
|
| 1.75e-153 | 634 | 1329 | 25 | 702 | Ascus wall endo-1,3(4)-beta-glucanase OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=eng2 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
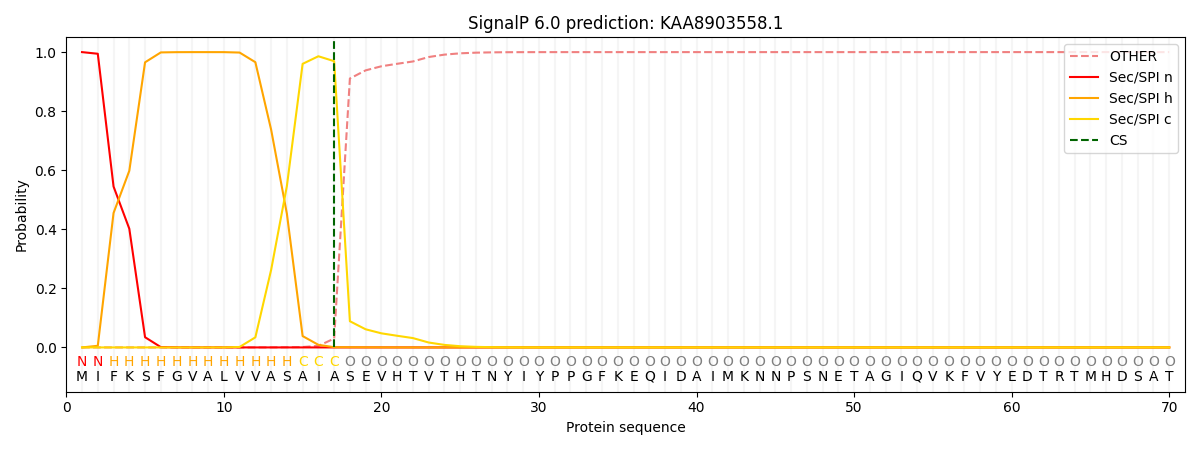
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000224 | 0.999738 | CS pos: 17-18. Pr: 0.9693 |
