You are browsing environment: FUNGIDB
CAZyme Information: K441DRAFT_71199-t45_1-p1
You are here: Home > Sequence: K441DRAFT_71199-t45_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Cenococcum geophilum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Dothideomycetes; ; Gloniaceae; Cenococcum; Cenococcum geophilum | |||||||||||
| CAZyme ID | K441DRAFT_71199-t45_1-p1 | |||||||||||
| CAZy Family | GT69 | |||||||||||
| CAZyme Description | carbohydrate-binding module family 18 protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.14:3 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH18 | 31 | 304 | 3.3e-22 | 0.6790540540540541 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 119356 | GH18_hevamine_XipI_class_III | 6.47e-93 | 31 | 344 | 1 | 271 | This conserved domain family includes xylanase inhibitor Xip-I, and the class III plant chitinases such as hevamine, concanavalin B, and PPL2, all of which have a glycosyl hydrolase family 18 (GH18) domain. Hevamine is a class III endochitinase that hydrolyzes the linear polysaccharide chains of chitin and peptidoglycan and is important for defense against pathogenic bacteria and fungi. PPL2 (Parkia platycephala lectin 2) is a class III chitinase from Parkia platycephala seeds that hydrolyzes beta(1-4) glycosidic bonds linking 2-acetoamido-2-deoxy-beta-D-glucopyranose units in chitin. |
| 395135 | Chitin_bind_1 | 6.79e-12 | 412 | 447 | 2 | 34 | Chitin recognition protein. |
| 395573 | Glyco_hydro_18 | 1.12e-11 | 32 | 297 | 1 | 222 | Glycosyl hydrolases family 18. |
| 211316 | ChtBD1_1 | 2.15e-11 | 412 | 448 | 2 | 37 | Hevein or type 1 chitin binding domain; filamentous ascomycete subfamily. Hevein or type 1 chitin binding domain (ChtBD1), a lectin domain found in proteins from plants and fungi that bind N-acetylglucosamine, plant endochitinases, wound-induced proteins such as hevein, a major IgE-binding allergen in natural rubber latex, and the alpha subunit of Kluyveromyces lactis killer toxin. This domain is involved in the recognition and/or binding of chitin subunits; it typically occurs N-terminal to glycosyl hydrolase domains in chitinases, together with other carbohydrate-binding domains, or by itself in tandem-repeat arrangements. |
| 211311 | ChtBD1 | 2.63e-08 | 412 | 447 | 2 | 34 | Hevein or type 1 chitin binding domain. Hevein or type 1 chitin binding domain (ChtBD1), a lectin domain found in proteins from plants and fungi that bind N-acetylglucosamine, plant endochitinases, wound-induced proteins such as hevein, a major IgE-binding allergen in natural rubber latex, and the alpha subunit of Kluyveromyces lactis killer toxin. This domain is involved in the recognition and/or binding of chitin subunits; it typically occurs N-terminal to glycosyl hydrolase domains in chitinases, together with other carbohydrate-binding domains, or by itself in tandem-repeat arrangements. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 4.16e-111 | 6 | 367 | 4 | 357 | |
| 5.15e-111 | 6 | 367 | 4 | 357 | |
| 7.27e-111 | 6 | 367 | 4 | 357 | |
| 3.32e-109 | 6 | 356 | 4 | 346 | |
| 1.03e-99 | 13 | 384 | 12 | 372 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 9.07e-91 | 30 | 360 | 1 | 309 | AfChiA1 in complex with compound 1 [Aspergillus fumigatus A1163],4TX6_B AfChiA1 in complex with compound 1 [Aspergillus fumigatus A1163] |
|
| 2.45e-90 | 31 | 360 | 1 | 308 | A. fumigatus chitinase A1 phenyl-methylguanylurea complex [Aspergillus fumigatus],2XVN_B A. fumigatus chitinase A1 phenyl-methylguanylurea complex [Aspergillus fumigatus],2XVN_C A. fumigatus chitinase A1 phenyl-methylguanylurea complex [Aspergillus fumigatus] |
|
| 2.53e-90 | 31 | 360 | 2 | 309 | ChiA1 from Aspergillus fumigatus in complex with acetazolamide [Aspergillus fumigatus A1163],2XTK_B ChiA1 from Aspergillus fumigatus in complex with acetazolamide [Aspergillus fumigatus A1163],2XUC_A Natural product-guided discovery of a fungal chitinase inhibitor [Aspergillus fumigatus],2XUC_B Natural product-guided discovery of a fungal chitinase inhibitor [Aspergillus fumigatus],2XUC_C Natural product-guided discovery of a fungal chitinase inhibitor [Aspergillus fumigatus],2XVP_A ChiA1 from Aspergillus fumigatus, apostructure [Aspergillus fumigatus A1163],2XVP_B ChiA1 from Aspergillus fumigatus, apostructure [Aspergillus fumigatus A1163] |
|
| 1.65e-54 | 26 | 362 | 1 | 290 | ScCTS1_apo crystal structure [Saccharomyces cerevisiae],2UY3_A ScCTS1_8-chlorotheophylline crystal structure [Saccharomyces cerevisiae],2UY4_A ScCTS1_acetazolamide crystal structure [Saccharomyces cerevisiae],2UY5_A ScCTS1_kinetin crystal structure [Saccharomyces cerevisiae],4TXE_A ScCTS1 in complex with compound 5 [Saccharomyces cerevisiae] |
|
| 1.93e-33 | 33 | 336 | 3 | 255 | CRYSTAL STRUCTURES OF HEVAMINE, A PLANT DEFENCE PROTEIN WITH CHITINASE AND LYSOZYME ACTIVITY, AND ITS COMPLEX WITH AN INHIBITOR [Hevea brasiliensis],1LLO_A Chain A, HEVAMINE [Hevea brasiliensis],2HVM_A Hevamine A At 1.8 Angstrom Resolution [Hevea brasiliensis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 8.63e-88 | 7 | 360 | 6 | 336 | Endochitinase A1 OS=Neosartorya fumigata OX=746128 GN=chiA1 PE=1 SV=1 |
|
| 1.00e-86 | 7 | 360 | 6 | 336 | Endochitinase A1 OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=chiA1 PE=3 SV=1 |
|
| 8.90e-86 | 20 | 360 | 18 | 337 | Endochitinase A OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=ctcA PE=2 SV=1 |
|
| 1.99e-80 | 18 | 361 | 16 | 338 | Endochitinase A OS=Emericella nidulans OX=162425 GN=chiA PE=2 SV=3 |
|
| 1.99e-80 | 18 | 361 | 16 | 338 | Endochitinase A OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=chiA PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
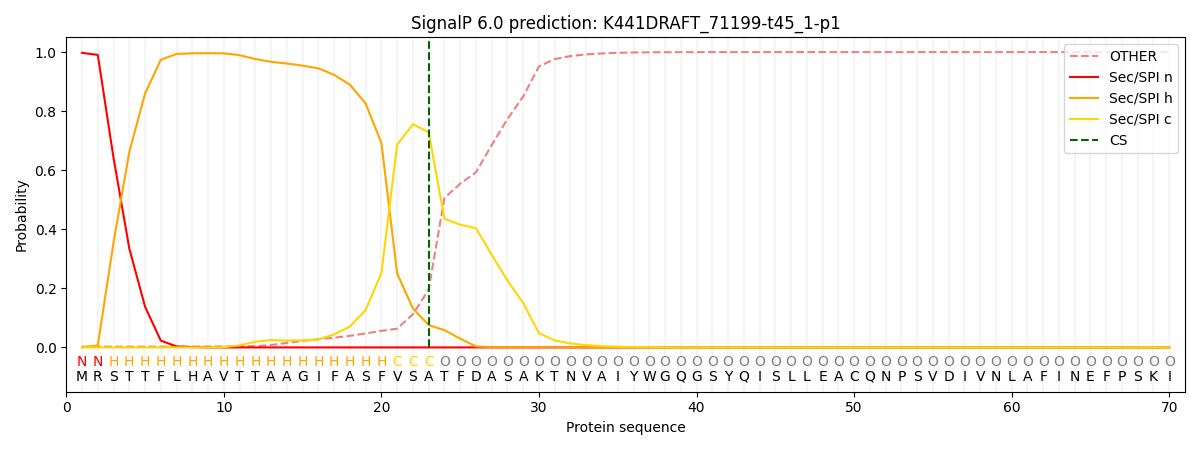
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.004519 | 0.995470 | CS pos: 23-24. Pr: 0.7282 |
