You are browsing environment: FUNGIDB
CAZyme Information: K441DRAFT_599654-t45_1-p1
You are here: Home > Sequence: K441DRAFT_599654-t45_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
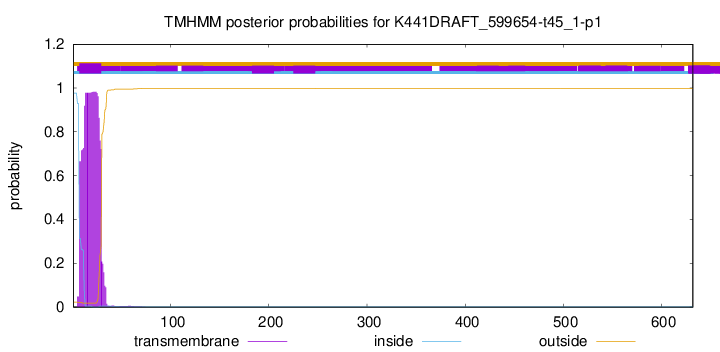
TMHMM annotations
Basic Information help
| Species | Cenococcum geophilum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Dothideomycetes; ; Gloniaceae; Cenococcum; Cenococcum geophilum | |||||||||||
| CAZyme ID | K441DRAFT_599654-t45_1-p1 | |||||||||||
| CAZy Family | CE16 | |||||||||||
| CAZyme Description | GMC oxidoreductase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| AA3 | 54 | 627 | 2.5e-186 | 0.9947183098591549 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 235000 | PRK02106 | 4.35e-112 | 54 | 626 | 5 | 532 | choline dehydrogenase; Validated |
| 273814 | betA | 1.36e-94 | 56 | 626 | 1 | 527 | choline dehydrogenase. Choline dehydrogenase catalyzes the conversion of exogenously supplied choline into the intermediate glycine betaine aldehyde, as part of a two-step oxidative reaction leading to the formation of osmoprotectant betaine. This enzymatic system can be found in both gram-positive and gram-negative bacteria. As in Escherichia coli, Staphylococcus xylosus, and Sinorhizobium meliloti, this enzyme is found associated in a transciptionally co-induced gene cluster with betaine aldehyde dehydrogenase, the second catalytic enzyme in this reaction. Other gram-positive organisms have been shown to employ a different enzymatic system, utlizing a soluable choline oxidase or type III alcohol dehydrogenase instead of choline dehydrogenase. This enzyme is a member of the GMC oxidoreductase family (pfam00732 and pfam05199), sharing a common evoluntionary origin and enzymatic reaction with alcohol dehydrogenase. Outgrouping from this model, Caulobacter crescentus shares sequence homology with choline dehydrogenase, yet other genes participating in this enzymatic reaction have not currently been identified. [Cellular processes, Adaptations to atypical conditions] |
| 225186 | BetA | 1.49e-92 | 52 | 629 | 5 | 537 | Choline dehydrogenase or related flavoprotein [Lipid transport and metabolism, General function prediction only]. |
| 398739 | GMC_oxred_C | 2.73e-44 | 482 | 621 | 1 | 143 | GMC oxidoreductase. This domain found associated with pfam00732. |
| 215420 | PLN02785 | 4.95e-30 | 42 | 614 | 43 | 565 | Protein HOTHEAD |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 2.13e-191 | 52 | 628 | 31 | 599 | |
| 2.12e-190 | 41 | 626 | 23 | 625 | |
| 6.73e-190 | 37 | 626 | 27 | 618 | |
| 2.62e-189 | 51 | 630 | 32 | 616 | |
| 2.25e-188 | 44 | 626 | 34 | 619 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5.43e-67 | 52 | 630 | 3 | 587 | Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE2_B Chain B, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE3_A Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE4_A Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE4_B Chain B, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE5_A Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE5_B Chain B, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE6_A Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE6_B Chain B, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE7_A Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE7_B Chain B, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],7AA2_A Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],7AA2_B Chain B, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495] |
|
| 1.11e-65 | 53 | 630 | 1 | 566 | Crystal structure of aryl-alcohol-oxidase from Pleurotus eryingii [Pleurotus eryngii] |
|
| 1.51e-65 | 55 | 630 | 2 | 565 | Crystal structure of aryl-alcohol oxidase from Pleurotus eryngii in complex with p-anisic acid [Pleurotus eryngii] |
|
| 6.65e-57 | 55 | 625 | 14 | 526 | Crystal structure of choline oxidase S101A mutant [Arthrobacter globiformis],3NNE_B Crystal structure of choline oxidase S101A mutant [Arthrobacter globiformis],3NNE_C Crystal structure of choline oxidase S101A mutant [Arthrobacter globiformis],3NNE_D Crystal structure of choline oxidase S101A mutant [Arthrobacter globiformis],3NNE_E Crystal structure of choline oxidase S101A mutant [Arthrobacter globiformis],3NNE_F Crystal structure of choline oxidase S101A mutant [Arthrobacter globiformis],3NNE_G Crystal structure of choline oxidase S101A mutant [Arthrobacter globiformis],3NNE_H Crystal structure of choline oxidase S101A mutant [Arthrobacter globiformis] |
|
| 1.36e-56 | 55 | 629 | 41 | 601 | Crystal structure of pyranose dehydrogenase from Agaricus meleagris, wildtype [Leucoagaricus meleagris] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 8.85e-179 | 41 | 625 | 38 | 624 | Patulin synthase OS=Aspergillus clavatus (strain ATCC 1007 / CBS 513.65 / DSM 816 / NCTC 3887 / NRRL 1 / QM 1276 / 107) OX=344612 GN=patE PE=1 SV=1 |
|
| 4.16e-174 | 51 | 625 | 48 | 624 | Patulin synthase OS=Penicillium expansum OX=27334 GN=patE PE=1 SV=1 |
|
| 3.34e-163 | 46 | 626 | 27 | 608 | Oxidoreductase cicC OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=cicC PE=1 SV=1 |
|
| 2.67e-156 | 41 | 628 | 60 | 642 | Versicolorin B synthase OS=Aspergillus parasiticus (strain ATCC 56775 / NRRL 5862 / SRRC 143 / SU-1) OX=1403190 GN=aflK PE=1 SV=1 |
|
| 3.12e-155 | 52 | 629 | 38 | 611 | Dehydrogenase pkfF OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=pkfF PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
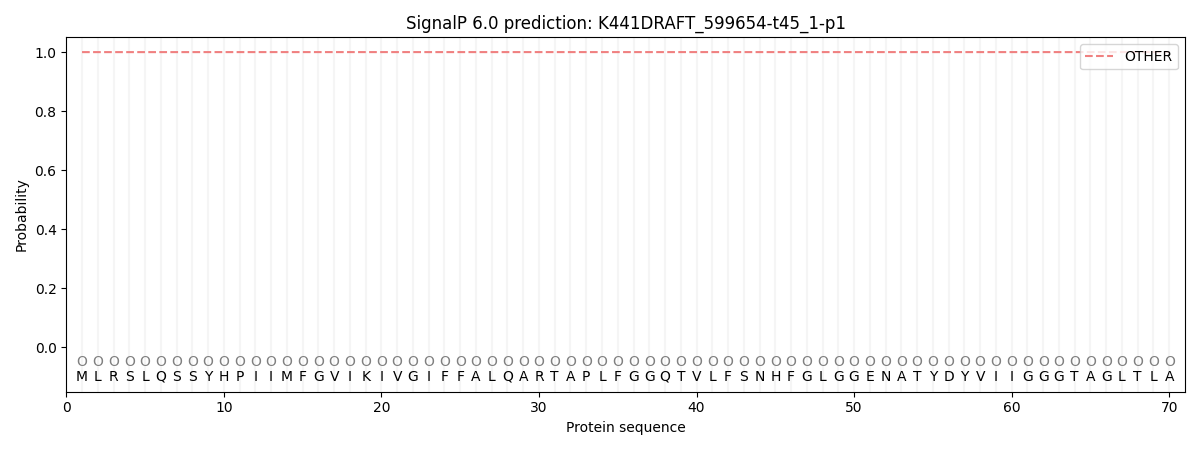
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.999964 | 0.000061 |