You are browsing environment: FUNGIDB
CAZyme Information: I317_03860-t43_1-p1
You are here: Home > Sequence: I317_03860-t43_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Kwoniella heveanensis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Basidiomycota; Tremellomycetes; ; Cryptococcaceae; Kwoniella; Kwoniella heveanensis | |||||||||||
| CAZyme ID | I317_03860-t43_1-p1 | |||||||||||
| CAZy Family | GH3 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT71 | 288 | 515 | 8.9e-69 | 0.9962121212121212 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 402574 | Mannosyl_trans3 | 1.13e-27 | 288 | 515 | 1 | 273 | Mannosyltransferase putative. This family is conserved in fungi. Several members are annotated as being alpha-1,3-mannosyltransferase but this could not be confirmed. |
| 133064 | GT8_GNT1 | 0.009 | 328 | 417 | 42 | 132 | GNT1 is a fungal enzyme that belongs to the GT 8 family. N-acetylglucosaminyltransferase is a fungal enzyme that catalyzes the addition of N-acetyl-D-glucosamine to mannotetraose side chains by an alpha 1-2 linkage during the synthesis of mannan. The N-acetyl-D-glucosamine moiety in mannan plays a role in the attachment of mannan to asparagine residues in proteins. The mannotetraose and its N-acetyl-D-glucosamine derivative side chains of mannan are the principle immunochemical determinants on the cell surface. N-acetylglucosaminyltransferase is a member of glycosyltransferase family 8, which are, based on the relative anomeric stereochemistry of the substrate and product in the reaction catalyzed, retaining glycosyltransferases. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 6 | 738 | 3 | 686 | |
| 0.0 | 6 | 738 | 3 | 686 | |
| 0.0 | 6 | 738 | 3 | 686 | |
| 0.0 | 6 | 738 | 3 | 679 | |
| 0.0 | 6 | 738 | 3 | 700 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.65e-13 | 219 | 504 | 186 | 519 | Alpha-1,2-mannosyltransferase MNN21 OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=MNN21 PE=3 SV=1 |
|
| 1.39e-09 | 283 | 509 | 181 | 459 | Alpha-1,2-mannosyltransferase MNN2 OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=MNN2 PE=1 SV=2 |
|
| 1.56e-09 | 287 | 474 | 291 | 518 | Alpha-1,2-mannosyltransferase MNN24 OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=MNN24 PE=3 SV=1 |
|
| 2.86e-08 | 290 | 513 | 164 | 434 | Alpha-1,2-mannosyltransferase MNN22 OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=MNN22 PE=2 SV=1 |
|
| 3.85e-08 | 285 | 413 | 169 | 302 | Alpha-1,2-mannosyltransferase MNN23 OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=MNN23 PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
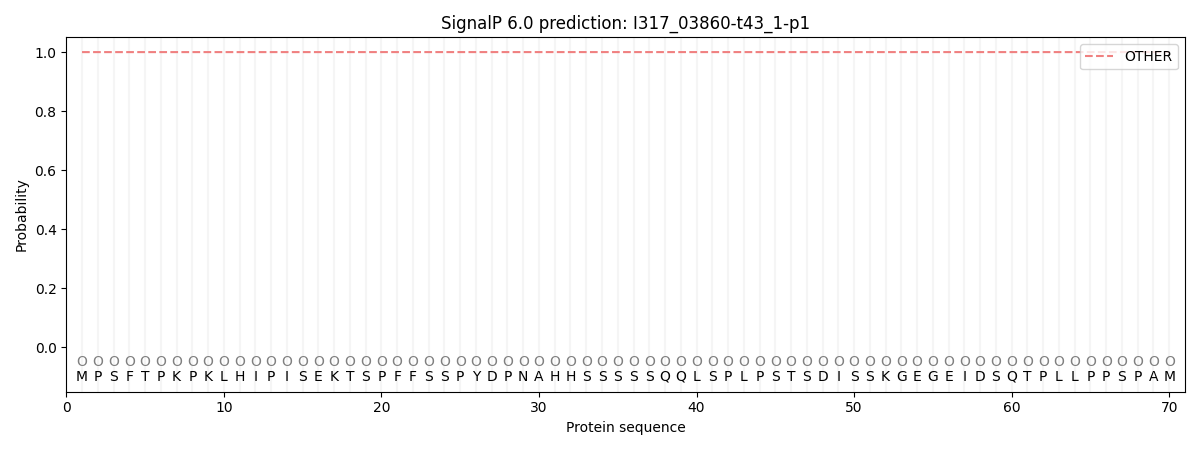
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000024 | 0.000000 |
