You are browsing environment: FUNGIDB
CAZyme Information: I311_04405-t41_1-p1
You are here: Home > Sequence: I311_04405-t41_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Cryptococcus gattii VGI | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Arthropoda; Insecta; ; Eriococcidae; Cryptococcus; Cryptococcus gattii VGI | |||||||||||
| CAZyme ID | I311_04405-t41_1-p1 | |||||||||||
| CAZy Family | PL14 | |||||||||||
| CAZyme Description | transglycosylase SLT domain-containing protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 17613; End:19079 Strand: - | |||||||||||
Full Sequence Download help
| MFANSLSLIT ALLPLLALTA QAAAAHSNQP PHLRHKRVLH HRRDTEHSHP HAARAQLPPR | 60 |
| SNAGHADAKK LIKKSVKKRG QTCRTRGSSY ATSSAYDVAA AQTLSSSSYV EPTSSSTAWS | 120 |
| DDSSSSSYAA AATPVAATLS NEQAWAPQVI SSSNDWSEST TADSWSSSAT SSSAWTQPTS | 180 |
| SFSSGSSSGS SSSGLLTITD AICGYSNADS EIPNGSEDWL NCGLNAAGWT PPMVTVDELI | 240 |
| ASELTTDGVF APCADYVDLF NQYADQYGLK GIMLASFAMQ ESTCNPSATG GNGEAGLMQL | 300 |
| ASENCGGAPN GNCYDVNFNI QRAAELFSNL ISSNGGNVLL AIGSYNGWYS GLTYAAGTAA | 360 |
| ASQGQCHAQN NLDYIHQFCN GWMQNKSGYT LGTYFNLKSC | 400 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 396169 | SLT | 1.92e-11 | 260 | 347 | 1 | 94 | Transglycosylase SLT domain. This family is distantly related to pfam00062. Members are found in phages, type II, type III and type IV secretion systems. |
| 381604 | Slt70-like | 3.91e-08 | 256 | 347 | 6 | 106 | 70kDa soluble lytic transglycosylase (Slt70) and similar proteins. Catalytic domain of the 70kda soluble lytic transglycosylase (LT)-like proteins, which also have an N-terminal U-shaped U-domain and a linker L-domain. LTs catalyze the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc), as do "goose-type" lysozymes. However, in addition to this, they also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. Proteins similar to this family include the soluble and insoluble membrane-bound LTs in bacteria and the LTs in bacteriophage lambda. |
| 223812 | MltE | 1.24e-07 | 255 | 364 | 138 | 256 | Soluble lytic murein transglycosylase and related regulatory proteins (some contain LysM/invasin domains) [Cell wall/membrane/envelope biogenesis]. |
| 381594 | LT-like | 4.16e-07 | 273 | 347 | 3 | 80 | lytic transglycosylase(LT)-like domain. Members include the soluble and insoluble membrane-bound LTs in bacteria and LTs in bacteriophage lambda. LTs catalyze the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc), as do "goose-type" lysozymes. However, in addition to this, they also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. |
| 381606 | MLTF-like | 1.38e-06 | 260 | 346 | 1 | 93 | membrane-bound lytic murein transglycosylase F (MLTF) and similar proteins. This subfamily includes membrane-bound lytic murein transglycosylase F (MltF, murein lyase F) that degrades murein glycan strands. It is responsible for catalyzing the release of 1,6-anhydromuropeptides from peptidoglycan. Lytic transglycosylase catalyzes the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc) as do goose-type lysozymes. However, in addition, it also makes a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ADV24528.1|GH23 | 2.62e-263 | 1 | 400 | 1 | 400 |
| KNX49777.2|GH23 | 5.00e-227 | 1 | 400 | 1 | 402 |
| AAW46053.1|GH23 | 3.42e-213 | 1 | 400 | 1 | 401 |
| AFR97355.1|GH23 | 1.66e-208 | 1 | 400 | 1 | 399 |
| AUB27369.1|GH23 | 1.66e-208 | 1 | 400 | 1 | 399 |
SignalP and Lipop Annotations help
This protein is predicted as SP
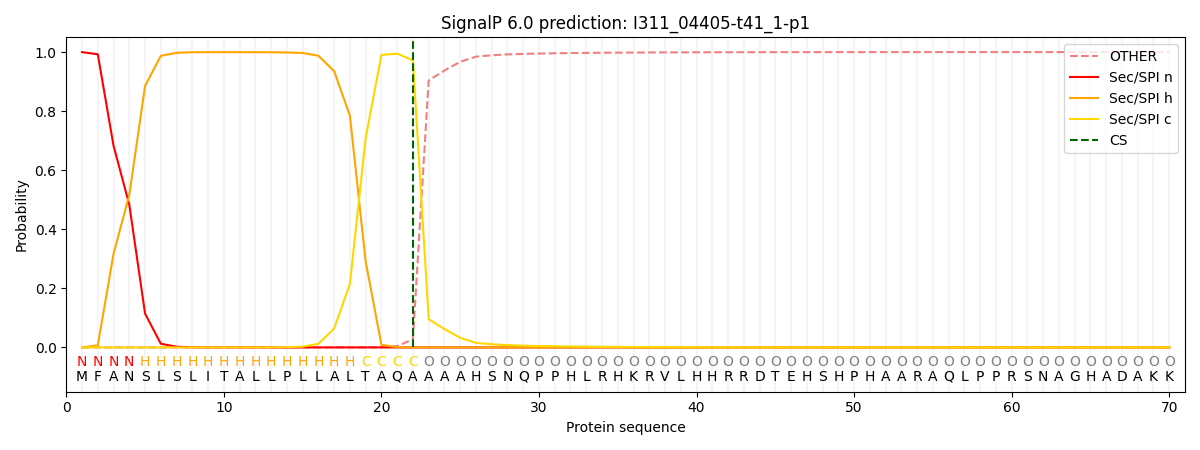
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000958 | 0.998993 | CS pos: 22-23. Pr: 0.9723 |
