You are browsing environment: FUNGIDB
CAZyme Information: I306_03435-t38_1-p1
You are here: Home > Sequence: I306_03435-t38_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
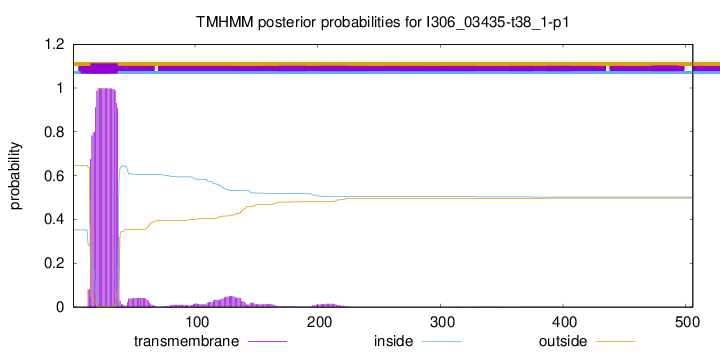
TMHMM annotations
Basic Information help
| Species | Cryptococcus gattii VGI | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Arthropoda; Insecta; ; Eriococcidae; Cryptococcus; Cryptococcus gattii VGI | |||||||||||
| CAZyme ID | I306_03435-t38_1-p1 | |||||||||||
| CAZy Family | GH37 | |||||||||||
| CAZyme Description | beta-1,4-mannosyltransferase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 2.4.1.142:10 | 2.4.1.-:1 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT33 | 49 | 497 | 6.6e-158 | 0.9905882352941177 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 340843 | GT33_ALG1-like | 0.0 | 48 | 500 | 3 | 411 | chitobiosyldiphosphodolichol beta-mannosyltransferase and similar proteins. This family is most closely related to the GT33 family of glycosyltransferases. The yeast gene ALG1 has been shown to function as a mannosyltransferase that catalyzes the formation of dolichol pyrophosphate (Dol-PP)-GlcNAc2Man from GDP-Man and Dol-PP-Glc-NAc2, and participates in the formation of the lipid-linked precursor oligosaccharide for N-glycosylation. In humans ALG1 has been associated with the congenital disorders of glycosylation (CDG) designated as subtype CDG-Ik. |
| 215155 | PLN02275 | 1.40e-132 | 49 | 447 | 5 | 371 | transferase, transferring glycosyl groups |
| 223515 | RfaB | 2.47e-08 | 90 | 473 | 25 | 367 | Glycosyltransferase involved in cell wall bisynthesis [Cell wall/membrane/envelope biogenesis]. |
| 340844 | GT4_UGDG-like | 2.22e-07 | 309 | 473 | 213 | 352 | UDP-Glc:1,2-diacylglycerol 3-a-glucosyltransferase and similar proteins. This family is most closely related to the GT1 family of glycosyltransferases. UDP-glucose-diacylglycerol glucosyltransferase (EC 2.4.1.337, UGDG; also known as 1,2-diacylglycerol 3-glucosyltransferase) catalyzes the transfer of glucose from UDP-glucose to 1,2-diacylglycerol forming 3-D-glucosyl-1,2-diacylglycerol. |
| 340831 | GT4_PimA-like | 4.77e-07 | 60 | 473 | 15 | 358 | phosphatidyl-myo-inositol mannosyltransferase. This family is most closely related to the GT4 family of glycosyltransferases and named after PimA in Propionibacterium freudenreichii, which is involved in the biosynthesis of phosphatidyl-myo-inositol mannosides (PIM) which are early precursors in the biosynthesis of lipomannans (LM) and lipoarabinomannans (LAM), and catalyzes the addition of a mannosyl residue from GDP-D-mannose (GDP-Man) to the position 2 of the carrier lipid phosphatidyl-myo-inositol (PI) to generate a phosphatidyl-myo-inositol bearing an alpha-1,2-linked mannose residue (PIM1). Glycosyltransferases catalyze the transfer of sugar moieties from activated donor molecules to specific acceptor molecules, forming glycosidic bonds. The acceptor molecule can be a lipid, a protein, a heterocyclic compound, or another carbohydrate residue. This group of glycosyltransferases is most closely related to the previously defined glycosyltransferase family 1 (GT1). The members of this family may transfer UDP, ADP, GDP, or CMP linked sugars. The diverse enzymatic activities among members of this family reflect a wide range of biological functions. The protein structure available for this family has the GTB topology, one of the two protein topologies observed for nucleotide-sugar-dependent glycosyltransferases. GTB proteins have distinct N- and C- terminal domains each containing a typical Rossmann fold. The two domains have high structural homology despite minimal sequence homology. The large cleft that separates the two domains includes the catalytic center and permits a high degree of flexibility. The members of this family are found mainly in certain bacteria and archaea. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 506 | 1 | 506 | |
| 0.0 | 1 | 506 | 1 | 506 | |
| 0.0 | 1 | 506 | 1 | 506 | |
| 0.0 | 1 | 506 | 1 | 506 | |
| 0.0 | 1 | 506 | 1 | 506 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.07e-102 | 49 | 496 | 33 | 457 | Chitobiosyldiphosphodolichol beta-mannosyltransferase OS=Pongo abelii OX=9601 GN=ALG1 PE=2 SV=1 |
|
| 2.31e-101 | 49 | 496 | 33 | 457 | Chitobiosyldiphosphodolichol beta-mannosyltransferase OS=Homo sapiens OX=9606 GN=ALG1 PE=1 SV=2 |
|
| 7.61e-98 | 49 | 496 | 33 | 457 | Chitobiosyldiphosphodolichol beta-mannosyltransferase OS=Mus musculus OX=10090 GN=Alg1 PE=1 SV=3 |
|
| 3.70e-90 | 51 | 451 | 7 | 421 | UDP-glycosyltransferase TURAN OS=Arabidopsis thaliana OX=3702 GN=TUN PE=2 SV=1 |
|
| 1.65e-81 | 53 | 488 | 6 | 455 | Chitobiosyldiphosphodolichol beta-mannosyltransferase OS=Dictyostelium discoideum OX=44689 GN=alg1 PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
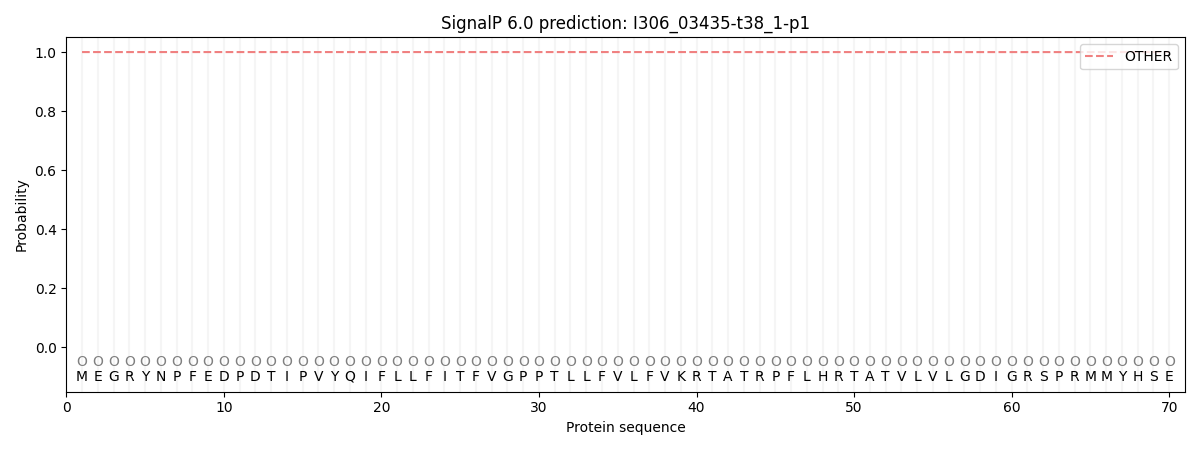
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000087 | 0.000000 |