You are browsing environment: FUNGIDB
CAZyme Information: HCBG_07743-t26_1-p1
You are here: Home > Sequence: HCBG_07743-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Histoplasma capsulatum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Ajellomycetaceae; Histoplasma; Histoplasma capsulatum | |||||||||||
| CAZyme ID | HCBG_07743-t26_1-p1 | |||||||||||
| CAZy Family | GT2|GT2 | |||||||||||
| CAZyme Description | choline dehydrogenase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| AA3 | 70 | 561 | 9.1e-139 | 0.8362676056338029 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 235000 | PRK02106 | 2.07e-60 | 44 | 567 | 4 | 538 | choline dehydrogenase; Validated |
| 225186 | BetA | 5.95e-56 | 17 | 564 | 50 | 537 | Choline dehydrogenase or related flavoprotein [Lipid transport and metabolism, General function prediction only]. |
| 398739 | GMC_oxred_C | 6.27e-35 | 414 | 556 | 1 | 143 | GMC oxidoreductase. This domain found associated with pfam00732. |
| 366272 | GMC_oxred_N | 3.80e-25 | 46 | 291 | 1 | 218 | GMC oxidoreductase. This family of proteins bind FAD as a cofactor. |
| 215420 | PLN02785 | 5.72e-12 | 23 | 549 | 38 | 565 | Protein HOTHEAD |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 71 | 568 | 1 | 498 | |
| 1.51e-285 | 148 | 558 | 1 | 411 | |
| 2.01e-183 | 35 | 564 | 19 | 614 | |
| 1.21e-143 | 45 | 563 | 29 | 614 | |
| 3.89e-142 | 37 | 564 | 30 | 623 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.51e-99 | 70 | 560 | 126 | 638 | Structural insights on a new fungal aryl-alcohol oxidase [Thermothelomyces thermophilus],6O9N_B Structural insights on a new fungal aryl-alcohol oxidase [Thermothelomyces thermophilus] |
|
| 4.85e-94 | 50 | 561 | 83 | 583 | Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE2_B Chain B, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE3_A Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE4_A Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE4_B Chain B, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE5_A Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE5_B Chain B, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE6_A Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE6_B Chain B, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE7_A Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],6ZE7_B Chain B, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],7AA2_A Chain A, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495],7AA2_B Chain B, FAD-dependent oxidoreductase [Thermochaetoides thermophila DSM 1495] |
|
| 1.14e-55 | 46 | 561 | 6 | 565 | Crystal structure of Aspergillus flavus FAD glucose dehydrogenase [Aspergillus flavus NRRL3357],4YNU_A Crystal structure of Aspergillus flavus FADGDH in complex with D-glucono-1,5-lactone [Aspergillus flavus NRRL3357] |
|
| 6.78e-46 | 46 | 561 | 17 | 588 | Chain A, Oligosaccharide dehydrogenase [Trametes cinnabarina],6XUU_A Chain A, Oligosaccharide dehydrogenase [Trametes cinnabarina],6XUV_A Chain A, Oligosaccharide dehydrogenase [Trametes cinnabarina] |
|
| 1.41e-43 | 65 | 561 | 86 | 562 | Crystal structure of aryl-alcohol oxidase from Pleurotus eryngii in complex with p-anisic acid [Pleurotus eryngii] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.37e-89 | 11 | 566 | 11 | 619 | Dehydrogenase xptC OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=xptC PE=3 SV=1 |
|
| 4.08e-61 | 18 | 562 | 13 | 617 | GMC oxidoreductase family protein Mala s 12 OS=Malassezia sympodialis (strain ATCC 42132) OX=1230383 GN=MSY001_2108 PE=1 SV=1 |
|
| 4.08e-61 | 18 | 562 | 13 | 617 | GMC oxidoreductase family protein Mala s 12.0101 OS=Malassezia sympodialis OX=76777 PE=1 SV=2 |
|
| 3.63e-55 | 18 | 562 | 9 | 597 | Dehydrogenase eriK OS=Hericium erinaceus OX=91752 GN=eriK PE=3 SV=1 |
|
| 6.42e-52 | 71 | 561 | 104 | 609 | Dehydrogenase mpl7 OS=Monascus purpureus OX=5098 GN=mpl7 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
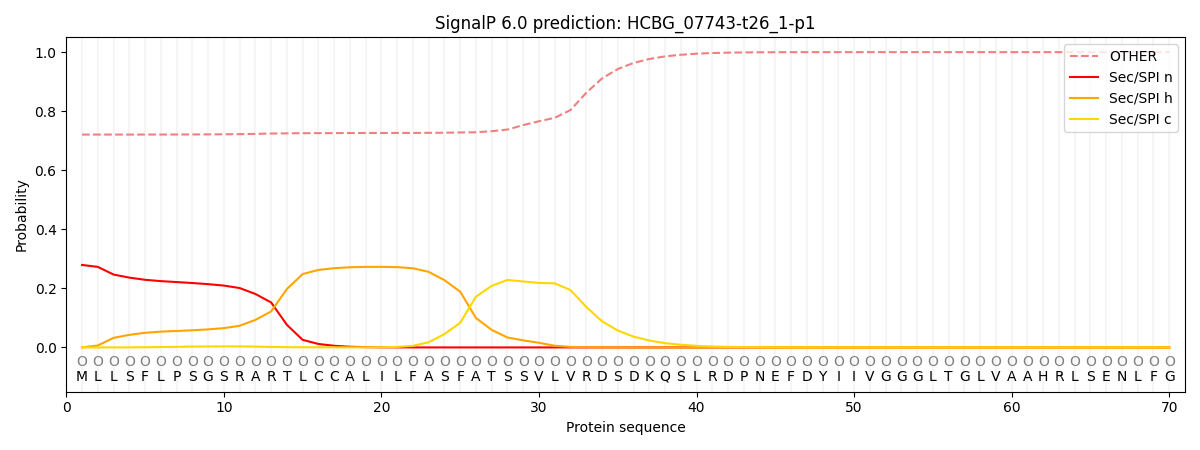
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.726998 | 0.272998 |
