You are browsing environment: FUNGIDB
CAZyme Information: H310_02571-t26_1-p1
You are here: Home > Sequence: H310_02571-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aphanomyces invadans | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Saprolegniaceae; Aphanomyces; Aphanomyces invadans | |||||||||||
| CAZyme ID | H310_02571-t26_1-p1 | |||||||||||
| CAZy Family | GH18 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.113:20 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH47 | 202 | 630 | 2.8e-134 | 0.9977578475336323 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 396217 | Glyco_hydro_47 | 5.77e-139 | 202 | 630 | 1 | 453 | Glycosyl hydrolase family 47. Members of this family are alpha-mannosidases that catalyze the hydrolysis of the terminal 1,2-linked alpha-D-mannose residues in the oligo-mannose oligosaccharide Man(9)(GlcNAc)(2). |
| 240427 | PTZ00470 | 6.53e-83 | 190 | 633 | 67 | 521 | glycoside hydrolase family 47 protein; Provisional |
| 400321 | PRKCSH | 6.12e-10 | 55 | 129 | 1 | 72 | Glucosidase II beta subunit-like protein. The sequences found in this family are similar to a region found in the beta-subunit of glucosidase II, which is also known as protein kinase C substrate 80K-H (PRKCSH). The enzyme catalyzes the sequential removal of two alpha-1,3-linked glucose residues in the second step of N-linked oligosaccharide processing. The beta subunit is required for the solubility and stability of the heterodimeric enzyme, and is involved in retaining the enzyme within the endoplasmic reticulum. Mutations in the gene coding for PRKCSH have been found to be involved in the development of autosomal dominant polycystic liver disease (ADPLD), but the precise role the protein has in the pathogenesis of this disease is unknown. This family also includes an ER sensor for misfolded glycoproteins and is therefore likely to be a generic sugar binding domain. |
| 404038 | PRKCSH_1 | 0.009 | 137 | 204 | 92 | 151 | Glucosidase II beta subunit-like protein. The sequences found in this family are similar to a region found in the beta-subunit of glucosidase II, which is also known as protein kinase C substrate 80K-H (PRKCSH). The enzyme catalyzes the sequential removal of two alpha-1,3-linked glucose residues in the second step of N-linked oligosaccharide processing. The beta subunit is required for the solubility and stability of the heterodimeric enzyme, and is involved in retaining the enzyme within the endoplasmic reticulum. The beta-subunit confers substrate specificity for di- and monoglucosylated glycans on the glucose-trimming activity of the alpha-subunit. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 55 | 1097 | 46 | 1072 | |
| 0.0 | 40 | 1055 | 19 | 1023 | |
| 2.30e-228 | 126 | 972 | 9 | 1035 | |
| 8.12e-129 | 187 | 632 | 32 | 475 | |
| 3.74e-127 | 199 | 632 | 50 | 481 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.05e-61 | 184 | 631 | 14 | 468 | Structure of mouse Golgi alpha-1,2-mannosidase IA and Man9GlcNAc2-PA complex [Mus musculus],5KKB_B Structure of mouse Golgi alpha-1,2-mannosidase IA and Man9GlcNAc2-PA complex [Mus musculus] |
|
| 5.03e-61 | 184 | 631 | 12 | 466 | Structure of mouse Golgi alpha-1,2-mannosidase IA reveals the molecular basis for substrate specificity among Class I enzymes (family 47 glycosidases) [Mus musculus] |
|
| 2.20e-53 | 185 | 626 | 3 | 431 | Structure of The GH47 processing alpha-1,2-mannosidase from Caulobacter strain K31 [Caulobacter sp. K31],4AYP_A Structure of The GH47 processing alpha-1,2-mannosidase from Caulobacter strain K31 in complex with thiomannobioside [Caulobacter sp. K31],4AYQ_A Structure of The GH47 processing alpha-1,2-mannosidase from Caulobacter strain K31 in complex with mannoimidazole [Caulobacter sp. K31],4AYR_A Structure of The GH47 processing alpha-1,2-mannosidase from Caulobacter strain K31 in complex with noeuromycin [Caulobacter sp. K31],5MEH_A Crystal structure of alpha-1,2-mannosidase from Caulobacter K31 strain in complex with 1-deoxymannojirimycin [Caulobacter sp. K31],5NE5_A Crystal structure of family 47 alpha-1,2-mannosidase from Caulobacter K31 strain in complex with kifunensine [Caulobacter sp. K31] |
|
| 1.14e-51 | 202 | 634 | 17 | 460 | Crystal Structure Of Human Class I Alpha1,2-Mannosidase In Complex With 1-Deoxymannojirimycin [Homo sapiens],1FO3_A Crystal Structure Of Human Class I Alpha1,2-Mannosidase In Complex With Kifunensine [Homo sapiens] |
|
| 2.33e-51 | 179 | 634 | 69 | 538 | Crystal Structure Of Human Class I alpha-1,2-Mannosidase In Complex With Thio-Disaccharide Substrate Analogue [Homo sapiens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.47e-126 | 193 | 632 | 38 | 475 | Alpha-mannosidase I MNS4 OS=Arabidopsis thaliana OX=3702 GN=MNS4 PE=1 SV=1 |
|
| 1.48e-121 | 184 | 631 | 25 | 479 | Alpha-mannosidase I MNS5 OS=Arabidopsis thaliana OX=3702 GN=MNS5 PE=1 SV=1 |
|
| 2.60e-117 | 199 | 646 | 39 | 495 | ER degradation-enhancing alpha-mannosidase-like protein 2 OS=Homo sapiens OX=9606 GN=EDEM2 PE=1 SV=2 |
|
| 1.89e-115 | 199 | 646 | 39 | 495 | ER degradation-enhancing alpha-mannosidase-like protein 2 OS=Mus musculus OX=10090 GN=Edem2 PE=1 SV=1 |
|
| 3.10e-102 | 192 | 626 | 122 | 577 | ER degradation-enhancing alpha-mannosidase-like protein 1 OS=Mus musculus OX=10090 GN=Edem1 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
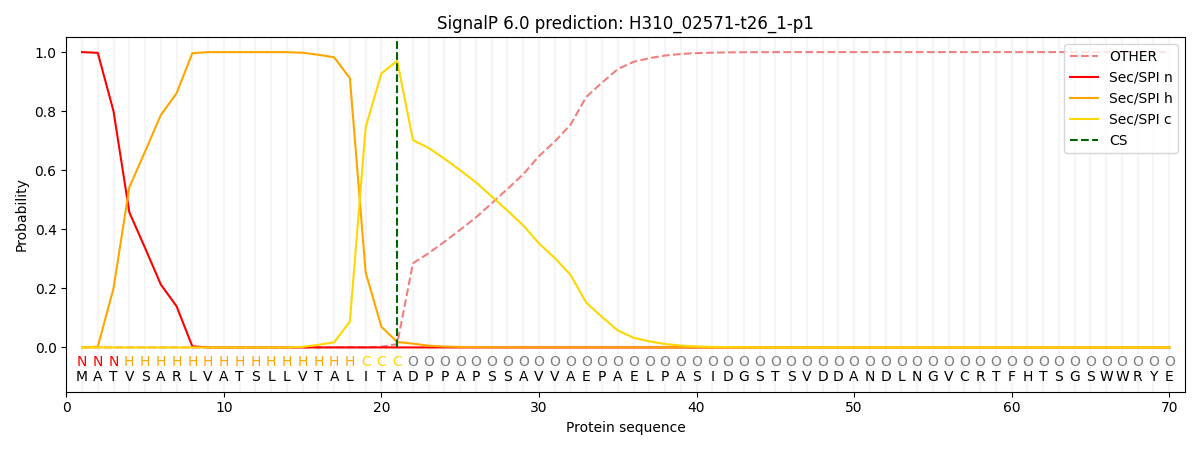
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000343 | 0.999629 | CS pos: 21-22. Pr: 0.9705 |
