You are browsing environment: FUNGIDB
CAZyme Information: H310_01082-t26_2-p1
You are here: Home > Sequence: H310_01082-t26_2-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aphanomyces invadans | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Saprolegniaceae; Aphanomyces; Aphanomyces invadans | |||||||||||
| CAZyme ID | H310_01082-t26_2-p1 | |||||||||||
| CAZy Family | GH16 | |||||||||||
| CAZyme Description | trehalose-phosphatase, variant | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT20 | 324 | 824 | 9.6e-128 | 0.9831578947368421 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 177855 | PLN02205 | 0.0 | 312 | 1142 | 43 | 848 | alpha,alpha-trehalose-phosphate synthase [UDP-forming] |
| 184712 | PRK14501 | 4.14e-167 | 330 | 1141 | 2 | 726 | putative bifunctional trehalose-6-phosphate synthase/HAD hydrolase subfamily IIB; Provisional |
| 340820 | GT20_TPS | 1.79e-143 | 331 | 823 | 2 | 463 | trehalose-6-phosphate synthase. Trehalose-6-Phosphate Synthase (TPS, EC 2.4.1.15) is a glycosyltransferase that catalyses the synthesis of alpha,alpha-1,1-trehalose-6-phosphate from glucose-6-phosphate using a UDP-glucose donor. It is a key enzyme in the trehalose synthesis pathway. Trehalose is a nonreducing disaccharide present in a wide variety of organisms and may serve as a source of energy and carbon. It is characterized most notably in insect, plant, and microbial cells. Its production is often associated with a variety of stress conditions, including desiccation, dehydration, heat, cold, and oxidation. This family represents the catalytic domain of the TPS. Some members of this domain family coexist with a C-terminal trehalose phosphatase domain. |
| 215555 | PLN03063 | 2.92e-143 | 331 | 1150 | 13 | 795 | alpha,alpha-trehalose-phosphate synthase (UDP-forming); Provisional |
| 215556 | PLN03064 | 2.33e-133 | 332 | 1107 | 97 | 815 | alpha,alpha-trehalose-phosphate synthase (UDP-forming); Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 1158 | 1 | 1096 | |
| 0.0 | 1 | 1158 | 1 | 1138 | |
| 2.91e-187 | 331 | 1155 | 24 | 846 | |
| 4.44e-181 | 120 | 1103 | 43 | 950 | |
| 1.78e-167 | 331 | 1142 | 57 | 836 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.22e-75 | 331 | 830 | 6 | 475 | Structure of Candida albicans trehalose-6-phosphate synthase in complex with UDP-glucose [Candida albicans SC5314],5HUT_B Structure of Candida albicans trehalose-6-phosphate synthase in complex with UDP-glucose [Candida albicans SC5314],5HUU_A Structure of Candida albicans trehalose-6-phosphate synthase in complex with UDP and glucose-6-phosphate [Candida albicans SC5314],5HUU_B Structure of Candida albicans trehalose-6-phosphate synthase in complex with UDP and glucose-6-phosphate [Candida albicans SC5314],5HVL_A Structure of Candida albicans trehalose-6-phosphate synthase in complex with UDP and validoxylamine A [Candida albicans SC5314],5HVL_B Structure of Candida albicans trehalose-6-phosphate synthase in complex with UDP and validoxylamine A [Candida albicans SC5314] |
|
| 5.79e-75 | 331 | 830 | 6 | 475 | Structure of Candida albicans trehalose-6-phosphate synthase E341R/E346R in complex with UDP-glucose [Candida albicans SC5314],5HUV_B Structure of Candida albicans trehalose-6-phosphate synthase E341R/E346R in complex with UDP-glucose [Candida albicans SC5314] |
|
| 1.44e-74 | 331 | 824 | 2 | 464 | Structure of Tps1 apo structure [Pyricularia oryzae 70-15],6JBI_B Structure of Tps1 apo structure [Pyricularia oryzae 70-15],6JBR_A Tps1/UDP/T6P complex [Pyricularia oryzae 70-15],6JBR_B Tps1/UDP/T6P complex [Pyricularia oryzae 70-15],6JBR_D Tps1/UDP/T6P complex [Pyricularia oryzae 70-15],6JBR_F Tps1/UDP/T6P complex [Pyricularia oryzae 70-15],6JBR_H Tps1/UDP/T6P complex [Pyricularia oryzae 70-15],6JBR_K Tps1/UDP/T6P complex [Pyricularia oryzae 70-15],6JBR_M Tps1/UDP/T6P complex [Pyricularia oryzae 70-15],6JBR_O Tps1/UDP/T6P complex [Pyricularia oryzae 70-15],6JBW_A Structure of Tps1/UDP complex [Pyricularia oryzae 70-15],6JBW_B Structure of Tps1/UDP complex [Pyricularia oryzae 70-15] |
|
| 2.86e-74 | 331 | 830 | 14 | 482 | Structure of Aspergillus fumigatus trehalose-6-phosphate synthase A in complex with UDP and validoxylamine A [Aspergillus fumigatus Af293],5HVM_B Structure of Aspergillus fumigatus trehalose-6-phosphate synthase A in complex with UDP and validoxylamine A [Aspergillus fumigatus Af293] |
|
| 1.39e-70 | 331 | 826 | 13 | 478 | Structure of Aspergillus fumigatus trehalose-6-phosphate synthase B in complex with UDP and validoxylamine A [Aspergillus fumigatus Af293],5HVO_B Structure of Aspergillus fumigatus trehalose-6-phosphate synthase B in complex with UDP and validoxylamine A [Aspergillus fumigatus Af293],5HVO_C Structure of Aspergillus fumigatus trehalose-6-phosphate synthase B in complex with UDP and validoxylamine A [Aspergillus fumigatus Af293],5HVO_D Structure of Aspergillus fumigatus trehalose-6-phosphate synthase B in complex with UDP and validoxylamine A [Aspergillus fumigatus Af293] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.56e-158 | 304 | 1144 | 39 | 845 | Alpha,alpha-trehalose-phosphate synthase [UDP-forming] 5 OS=Arabidopsis thaliana OX=3702 GN=TPS5 PE=1 SV=2 |
|
| 6.79e-155 | 325 | 1142 | 49 | 854 | Alpha,alpha-trehalose-phosphate synthase [UDP-forming] 6 OS=Arabidopsis thaliana OX=3702 GN=TPS6 PE=1 SV=2 |
|
| 2.03e-154 | 318 | 1154 | 48 | 847 | Probable alpha,alpha-trehalose-phosphate synthase [UDP-forming] 7 OS=Arabidopsis thaliana OX=3702 GN=TPS7 PE=1 SV=1 |
|
| 2.27e-154 | 332 | 1145 | 62 | 846 | Probable alpha,alpha-trehalose-phosphate synthase [UDP-forming] 9 OS=Arabidopsis thaliana OX=3702 GN=TPS9 PE=2 SV=1 |
|
| 1.57e-149 | 306 | 1138 | 42 | 834 | Probable alpha,alpha-trehalose-phosphate synthase [UDP-forming] 8 OS=Arabidopsis thaliana OX=3702 GN=TPS8 PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
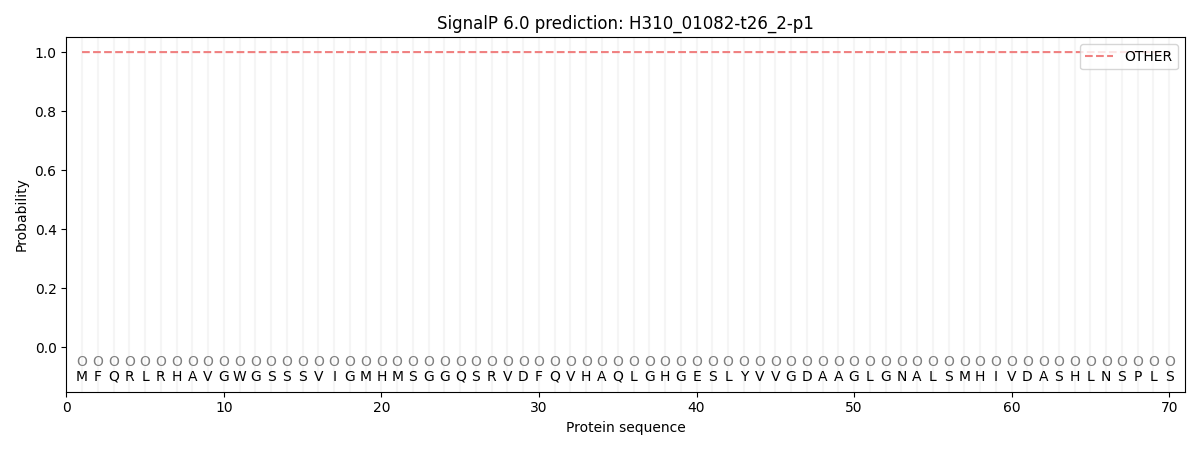
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000038 | 0.000006 |
