You are browsing environment: FUNGIDB
CAZyme Information: H257_12486-t26_1-p1
You are here: Home > Sequence: H257_12486-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aphanomyces astaci | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Saprolegniaceae; Aphanomyces; Aphanomyces astaci | |||||||||||
| CAZyme ID | H257_12486-t26_1-p1 | |||||||||||
| CAZy Family | GT20 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH72 | 21 | 325 | 2e-76 | 0.9006410256410257 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 397351 | Glyco_hydro_72 | 8.04e-56 | 19 | 378 | 5 | 315 | Glucanosyltransferase. This is a family of glycosylphosphatidylinositol-anchored beta(1-3)glucanosyltransferases. The active site residues in the Aspergillus fumigatus example are the two glutamate residues at 160 and 261. |
| 398978 | Epiglycanin_TR | 8.72e-04 | 508 | 563 | 10 | 65 | Tandem-repeating region of mucin, epiglycanin-like. The unusual mucin, epiglycanin, is membrane-bound at the C-terminus but has a long region of this tandem-repeat at the N-terminus. It was the first mucin identified to be associated with the malignant behaviour of carcinoma cells. Mouse Muc21/epiglycanin is thought to be a highly glycosylated molecule, which makes it likely that its function is dependent on its glycoforms. Cells expressing Muc21 are significantly less adherent to each other and to extracellular matrix components than control cells, and this loss of adhesion is mediated by the TR portion of Muc21. This family also now contains the repeat that was the C. elegans protein of unknown function (DUF801). |
| 398978 | Epiglycanin_TR | 0.005 | 508 | 568 | 3 | 58 | Tandem-repeating region of mucin, epiglycanin-like. The unusual mucin, epiglycanin, is membrane-bound at the C-terminus but has a long region of this tandem-repeat at the N-terminus. It was the first mucin identified to be associated with the malignant behaviour of carcinoma cells. Mouse Muc21/epiglycanin is thought to be a highly glycosylated molecule, which makes it likely that its function is dependent on its glycoforms. Cells expressing Muc21 are significantly less adherent to each other and to extracellular matrix components than control cells, and this loss of adhesion is mediated by the TR portion of Muc21. This family also now contains the repeat that was the C. elegans protein of unknown function (DUF801). |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 2.46e-211 | 19 | 439 | 34 | 451 | |
| 1.33e-209 | 11 | 437 | 3 | 445 | |
| 4.32e-194 | 19 | 447 | 16 | 447 | |
| 8.94e-169 | 25 | 448 | 24 | 451 | |
| 8.69e-165 | 11 | 433 | 4 | 429 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.68e-29 | 17 | 399 | 27 | 368 | Saccharomyces cerevisiae Gas2p in complex with laminaripentaose [Saccharomyces cerevisiae],2W63_A Saccharomyces Cerevisiae Gas2p In Complex With Laminaritriose And Laminaritetraose [Saccharomyces cerevisiae],5O9O_A Crystal structure of ScGas2 in complex with compound 7. [Saccharomyces cerevisiae S288C],5O9P_A Crystal structure of Gas2 in complex with compound 10 [Saccharomyces cerevisiae S288C],5O9Q_A Crystal structure of ScGas2 in complex with compound 6 [Saccharomyces cerevisiae S288C],5O9R_A Crystal structure of ScGas2 in complex with compound 9 [Saccharomyces cerevisiae S288C],5O9Y_A Crystal structure of ScGas2 in complex with compound 11 [Saccharomyces cerevisiae S288C],5OA2_A Crystal structure of ScGas2 in complex with compound 8 [Saccharomyces cerevisiae S288C],5OA2_B Crystal structure of ScGas2 in complex with compound 8 [Saccharomyces cerevisiae S288C],5OA2_C Crystal structure of ScGas2 in complex with compound 8 [Saccharomyces cerevisiae S288C],5OA6_A Crystal structure of ScGas2 in complex with compound 12 [Saccharomyces cerevisiae S288C] |
|
| 6.51e-29 | 17 | 399 | 27 | 368 | Saccharomyces cerevisiae Gas2p apostructure (E176Q mutant) [Saccharomyces cerevisiae] |
|
| 6.51e-29 | 17 | 399 | 27 | 368 | SACCHAROMYCES CEREVISIAE GAS2P (E176Q MUTANT) IN COMPLEX WITH LAMINARITETRAOSE AND LAMINARIPENTAOSE [Saccharomyces cerevisiae] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.71e-34 | 25 | 403 | 27 | 371 | 1,3-beta-glucanosyltransferase gas4 OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=gas4 PE=1 SV=1 |
|
| 8.11e-34 | 12 | 325 | 6 | 294 | 1,3-beta-glucanosyltransferase PGA4 OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=PGA4 PE=1 SV=1 |
|
| 6.09e-33 | 21 | 391 | 24 | 347 | 1,3-beta-glucanosyltransferase gas5 OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=gas5 PE=1 SV=1 |
|
| 3.33e-32 | 21 | 440 | 27 | 383 | 1,3-beta-glucanosyltransferase gel1 OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=gel1 PE=3 SV=1 |
|
| 4.53e-32 | 21 | 440 | 27 | 383 | 1,3-beta-glucanosyltransferase gel1 OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=gel1 PE=1 SV=1 |
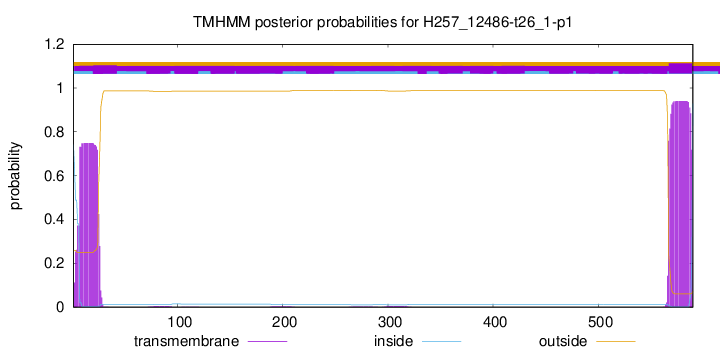
SignalP and Lipop Annotations help
This protein is predicted as SP
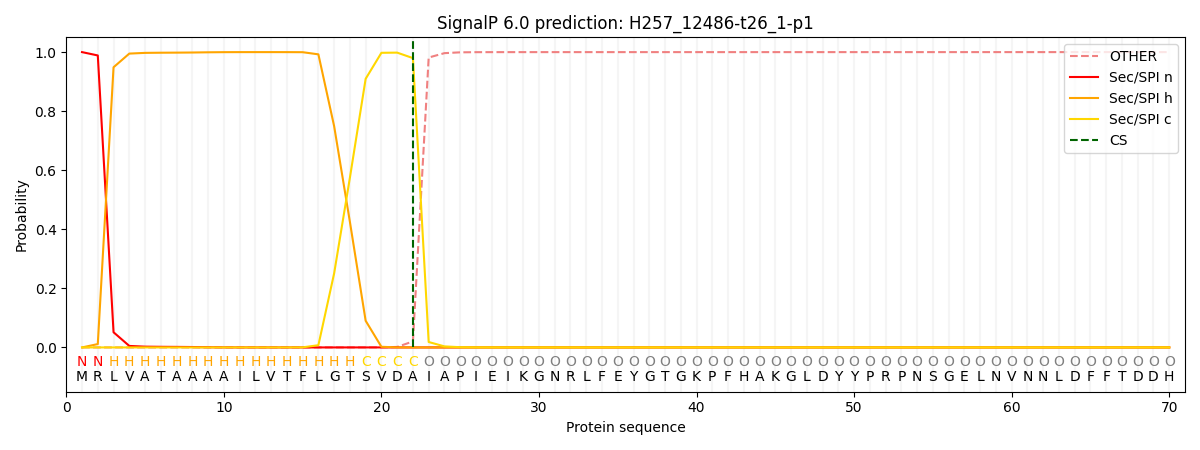
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000226 | 0.999735 | CS pos: 22-23. Pr: 0.9796 |