You are browsing environment: FUNGIDB
CAZyme Information: H257_04761-t26_2-p1
You are here: Home > Sequence: H257_04761-t26_2-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aphanomyces astaci | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Saprolegniaceae; Aphanomyces; Aphanomyces astaci | |||||||||||
| CAZyme ID | H257_04761-t26_2-p1 | |||||||||||
| CAZy Family | GH18 | |||||||||||
| CAZyme Description | hypothetical protein, variant 1 | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT60 | 348 | 535 | 3.2e-56 | 0.6454545454545455 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 371506 | GlcNAc | 3.49e-42 | 348 | 535 | 1 | 231 | Glycosyltransferase (GlcNAc). GlcNAc is an enzyme that carries out the first glycosylation step of hydroxylated Skp1, a ubiquitous eukaryotic protein, in the cytoplasm. |
| 404519 | 2OG-FeII_Oxy_3 | 4.63e-22 | 197 | 293 | 2 | 94 | 2OG-Fe(II) oxygenase superfamily. This family contains members of the 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily. |
| 214780 | P4Hc | 1.14e-14 | 134 | 292 | 20 | 164 | Prolyl 4-hydroxylase alpha subunit homologues. Mammalian enzymes catalyse hydroxylation of collagen, for example. Prokaryotic enzymes might catalyse hydroxylation of antibiotic peptides. These are 2-oxoglutarate-dependent dioxygenases, requiring 2-oxoglutarate and dioxygen as cosubstrates and ferrous iron as a cofactor. |
| 379319 | 2OG-FeII_Oxy_4 | 2.21e-10 | 226 | 292 | 22 | 92 | 2OG-Fe(II) oxygenase superfamily. This family contains members of the 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily. |
| 226274 | EGL9 | 1.18e-09 | 146 | 292 | 88 | 238 | Proline 4-hydroxylase (includes Rps23 Pro-64 3,4-dihydroxylase Tpa1), contains SM-20 domain [Translation, ribosomal structure and biogenesis, Posttranslational modification, protein turnover, chaperones]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 2.67e-84 | 14 | 534 | 37 | 518 | |
| 4.51e-47 | 149 | 535 | 107 | 477 | |
| 1.76e-43 | 91 | 534 | 62 | 493 | |
| 2.07e-42 | 347 | 535 | 4 | 209 | |
| 9.08e-30 | 346 | 535 | 2 | 193 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.03e-15 | 99 | 297 | 43 | 226 | Crystal structure of a Pseudomonas putida prolyl-4-hydroxylase (P4H) in complex with elongation factor Tu (EF-Tu) [Pseudomonas putida KT2440],4IW3_J Crystal structure of a Pseudomonas putida prolyl-4-hydroxylase (P4H) in complex with elongation factor Tu (EF-Tu) [Pseudomonas putida KT2440],4J25_A Crystal structure of a Pseudomonas putida prolyl-4-hydroxylase (P4H) [Pseudomonas putida KT2440],4J25_B Crystal structure of a Pseudomonas putida prolyl-4-hydroxylase (P4H) [Pseudomonas putida KT2440],4J25_C Crystal structure of a Pseudomonas putida prolyl-4-hydroxylase (P4H) [Pseudomonas putida KT2440],4J25_D Crystal structure of a Pseudomonas putida prolyl-4-hydroxylase (P4H) [Pseudomonas putida KT2440],4J25_E Crystal structure of a Pseudomonas putida prolyl-4-hydroxylase (P4H) [Pseudomonas putida KT2440],4J25_F Crystal structure of a Pseudomonas putida prolyl-4-hydroxylase (P4H) [Pseudomonas putida KT2440],4J25_G Crystal structure of a Pseudomonas putida prolyl-4-hydroxylase (P4H) [Pseudomonas putida KT2440],4J25_H Crystal structure of a Pseudomonas putida prolyl-4-hydroxylase (P4H) [Pseudomonas putida KT2440] |
|
| 9.69e-13 | 91 | 293 | 53 | 244 | prolyl hydroxylase from Trichoplax adhaerens [Trichoplax adhaerens] |
|
| 9.69e-13 | 91 | 293 | 53 | 244 | prolyl hydroxylase in complex with hypoxia inducible factor oxygen degradation domain peptide fragment from Trichoplax adhaerens [Trichoplax adhaerens] |
|
| 1.63e-12 | 99 | 293 | 27 | 212 | Hypoxia-Inducible Factor (HIF) Prolyl Hydroxylase 2 (PHD2) in Complex with the Carboxamide Analog JNJ43058171 [Homo sapiens] |
|
| 1.63e-12 | 99 | 293 | 27 | 212 | PHD2-R717 with 40787422 [Homo sapiens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.66e-43 | 347 | 535 | 4 | 209 | [Skp1-protein]-hydroxyproline N-acetylglucosaminyltransferase OS=Dictyostelium discoideum OX=44689 GN=gnt1 PE=1 SV=2 |
|
| 7.14e-17 | 99 | 293 | 27 | 213 | Prolyl hydroxylase EGLN3 OS=Mus musculus OX=10090 GN=Egln3 PE=1 SV=1 |
|
| 1.78e-16 | 99 | 293 | 27 | 213 | Prolyl hydroxylase EGLN3 OS=Homo sapiens OX=9606 GN=EGLN3 PE=1 SV=1 |
|
| 4.42e-16 | 99 | 293 | 27 | 213 | Prolyl hydroxylase EGLN3 OS=Rattus norvegicus OX=10116 GN=Egln3 PE=1 SV=2 |
|
| 6.56e-13 | 91 | 292 | 367 | 564 | Hypoxia-inducible factor prolyl hydroxylase OS=Caenorhabditis elegans OX=6239 GN=egl-9 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
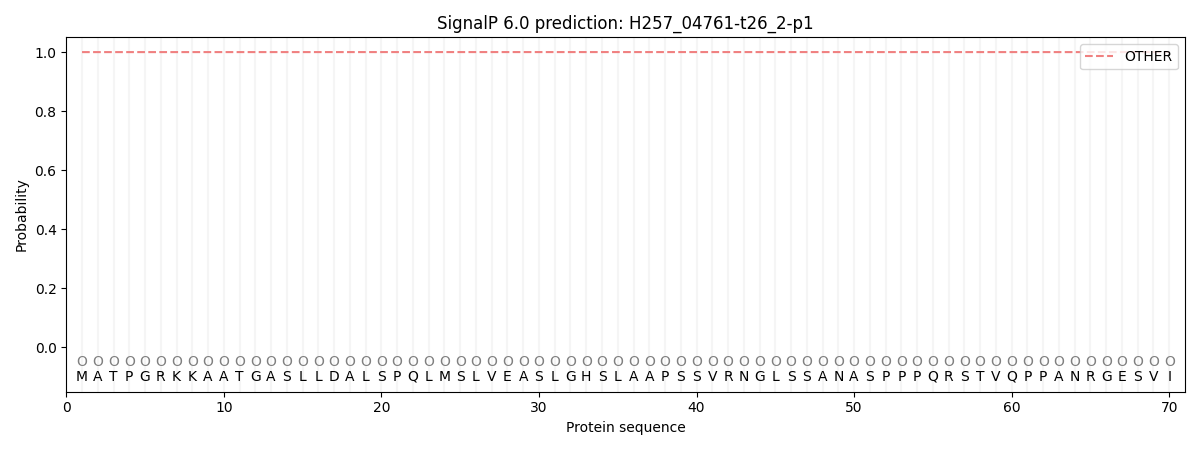
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000053 | 0.000000 |
