You are browsing environment: FUNGIDB
CAZyme Information: H257_01053-t26_1-p1
You are here: Home > Sequence: H257_01053-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aphanomyces astaci | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Saprolegniaceae; Aphanomyces; Aphanomyces astaci | |||||||||||
| CAZyme ID | H257_01053-t26_1-p1 | |||||||||||
| CAZy Family | AA15 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.113:20 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH47 | 38 | 482 | 4.3e-131 | 0.9977578475336323 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 396217 | Glyco_hydro_47 | 2.08e-137 | 38 | 482 | 1 | 453 | Glycosyl hydrolase family 47. Members of this family are alpha-mannosidases that catalyze the hydrolysis of the terminal 1,2-linked alpha-D-mannose residues in the oligo-mannose oligosaccharide Man(9)(GlcNAc)(2). |
| 240427 | PTZ00470 | 1.19e-78 | 33 | 484 | 74 | 520 | glycoside hydrolase family 47 protein; Provisional |
| 239041 | PA_EDEM3_like | 3.07e-17 | 540 | 666 | 2 | 126 | PA_EDEM3_like: protease associated domain (PA) domain-containing EDEM3-like proteins. This group contains various PA domain-containing proteins similar to mouse EDEM3 (ER-degradation-enhancing mannosidase-like 3 protein). EDEM3 contains a region, similar to Class I alpha-mannosidases (gylcosyl hydrolase family 47), N-terminal to the PA domain. EDEM3 accelerates glycoprotein ERAD (ER-associated degradation). In transfected mammalian cells, overexpression of EDEM3 enhances the mannose trimming from the N-glycans, of a model misfolded protein [alpha1-antitrypsin null (Hong Kong)] as well as, from total glycoproteins. Mannose trimming appears to be involved in the selection of ERAD substrates. EDEM3 has a different specificity of trimming than ER alpha-mannosidase 1. The significance of the PA domain to EDEM3 has not been ascertained. It may be a protein-protein interaction domain. At peptidase active sites, the PA domain may participate in substrate binding and/or promoting conformational changes, which influence the stability and accessibility of the site to substrate. |
| 239038 | PA_C_RZF_like | 4.71e-17 | 523 | 664 | 9 | 146 | PA_C-RZF_ like: Protease-associated (PA) domain C_RZF-like. This group includes various PA domain-containing proteins similar to C-RZF (chicken embryo RING zinc finger) protein. These proteins contain a C3H2C3 RING finger. C-RZF is expressed in embryo cells and is restricted mainly to brain and heart, it is localized to both the nucleus and endosomes. Additional C3H2C3 RING finger proteins belonging to this group, include Arabidopsis ReMembR-H2 protein and mouse sperizin. ReMembR-H2 is likely to be an integral membrane protein, and to traffic through the endosomal pathway. Sperizin is expressed in haploid germ cells and localized in the cytoplasm, it may participate in spermatogenesis. The significance of the PA domain to these proteins has not been ascertained. It may be a protein-protein interaction domain. At peptidase active sites, the PA domain may participate in substrate binding and/or promoting conformational changes, which influence the stability and accessibility of the site to substrate. |
| 240122 | PA_subtilisin_1 | 3.26e-13 | 542 | 651 | 2 | 104 | PA_subtilisin_1: Protease-associated domain containing subtilisin-like proteases, subgroup 1. A subgroup of PA domain-containing subtilisin-like proteases. The significance of the PA domain to many of the proteins in which it is inserted is undetermined. It may be a protein-protein interaction domain. At peptidase active sites, the PA domain may participate in substrate binding and/or promoting conformational changes, which influence the stability and accessibility of the site to substrate. Proteins into which the PA domain is inserted include the following subtilisin-like proteases: i) melon cucumisin, ii) Arabidopsis thaliana Ara12, iii) Alnus glutinosa ag12, iv) members of the tomato P69 family, and v) tomato LeSBT2. However, these proteins belong to other subtilisin-like subgroups. Relatively little is known about proteins in this subgroup. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 5.65e-303 | 13 | 670 | 14 | 696 | |
| 1.82e-216 | 24 | 647 | 23 | 681 | |
| 1.12e-141 | 38 | 510 | 1 | 461 | |
| 7.70e-131 | 27 | 486 | 31 | 472 | |
| 1.63e-130 | 6 | 486 | 19 | 481 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 9.85e-51 | 30 | 482 | 1 | 451 | Crystal structure of the class I human endoplasmic reticulum 1,2-alpha-mannosidase and Man9GlcNAc2-PA complex [Homo sapiens] |
|
| 1.12e-50 | 30 | 482 | 6 | 456 | Crystal Structure Of Human Class I Alpha1,2-Mannosidase [Homo sapiens] |
|
| 1.17e-50 | 30 | 482 | 6 | 456 | Crystal Structure Of Human Class I Alpha1,2-Mannosidase In Complex With 1-Deoxymannojirimycin [Homo sapiens],1FO3_A Crystal Structure Of Human Class I Alpha1,2-Mannosidase In Complex With Kifunensine [Homo sapiens] |
|
| 5.24e-50 | 30 | 482 | 1 | 451 | Crystal structure of the class I human endoplasmic reticulum 1,2-alpha-mannosidase T688A mutant and Thio-disaccharide substrate analog complex [Homo sapiens],5KK7_B Crystal structure of the class I human endoplasmic reticulum 1,2-alpha-mannosidase T688A mutant and Thio-disaccharide substrate analog complex [Homo sapiens] |
|
| 5.35e-50 | 30 | 482 | 84 | 534 | Crystal Structure Of Human Class I alpha-1,2-Mannosidase In Complex With Thio-Disaccharide Substrate Analogue [Homo sapiens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.75e-127 | 17 | 486 | 26 | 477 | Alpha-mannosidase I MNS4 OS=Arabidopsis thaliana OX=3702 GN=MNS4 PE=1 SV=1 |
|
| 2.55e-125 | 2 | 648 | 9 | 759 | ER degradation-enhancing alpha-mannosidase-like protein 3 OS=Xenopus laevis OX=8355 GN=edem3 PE=2 SV=2 |
|
| 2.41e-124 | 31 | 482 | 130 | 586 | ER degradation-enhancing alpha-mannosidase-like protein 1 OS=Homo sapiens OX=9606 GN=EDEM1 PE=1 SV=1 |
|
| 5.84e-124 | 31 | 482 | 125 | 581 | ER degradation-enhancing alpha-mannosidase-like protein 1 OS=Mus musculus OX=10090 GN=Edem1 PE=1 SV=1 |
|
| 8.09e-124 | 3 | 648 | 19 | 773 | ER degradation-enhancing alpha-mannosidase-like protein 3 OS=Mus musculus OX=10090 GN=Edem3 PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
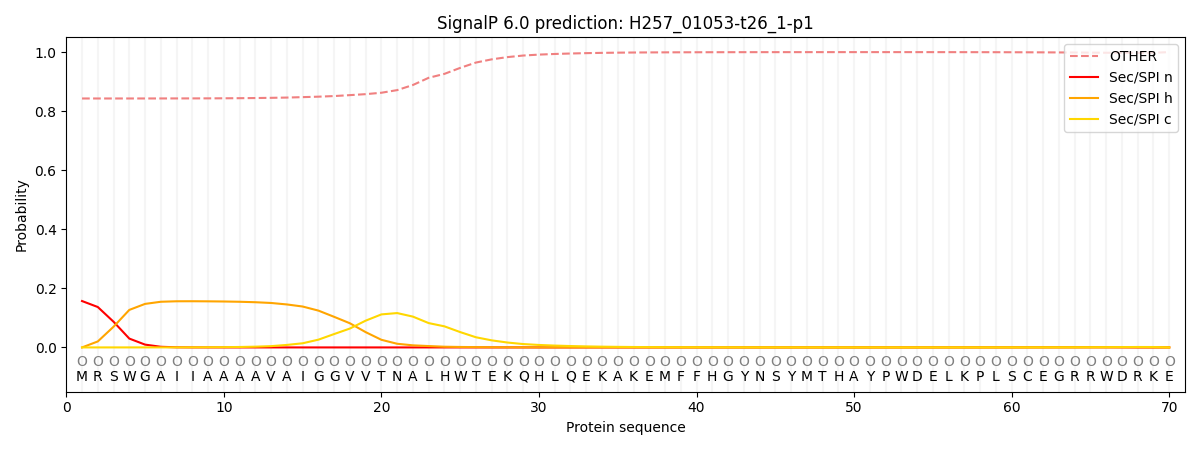
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.853819 | 0.146193 |

