You are browsing environment: FUNGIDB
CAZyme Information: GAQ09536.1
You are here: Home > Sequence: GAQ09536.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
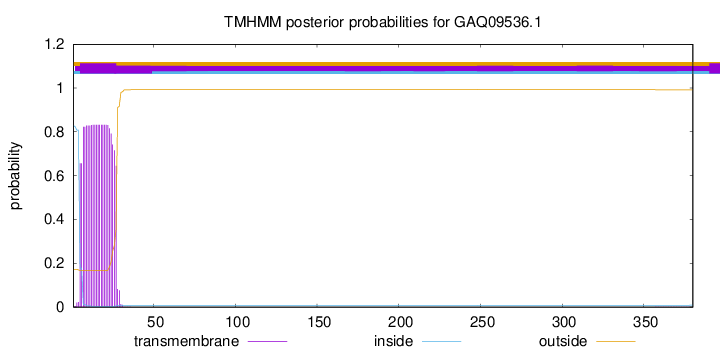
TMHMM annotations
Basic Information help
| Species | Aspergillus lentulus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus lentulus | |||||||||||
| CAZyme ID | GAQ09536.1 | |||||||||||
| CAZy Family | GH5 | |||||||||||
| CAZyme Description | probable pectin lyase A | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 4.2.2.10:25 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL1 | 114 | 298 | 1.3e-89 | 0.9893048128342246 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 214765 | Amb_all | 1.23e-55 | 114 | 300 | 2 | 190 | Amb_all domain. |
| 226384 | PelB | 3.89e-14 | 124 | 298 | 97 | 275 | Pectate lyase [Carbohydrate transport and metabolism]. |
| 366158 | Pec_lyase_C | 9.32e-13 | 124 | 296 | 30 | 211 | Pectate lyase. This enzyme forms a right handed beta helix structure. Pectate lyase is an enzyme involved in the maceration and soft rotting of plant tissue. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 4.17e-219 | 1 | 380 | 1 | 376 | |
| 1.97e-217 | 1 | 380 | 1 | 376 | |
| 6.01e-215 | 1 | 377 | 1 | 377 | |
| 5.11e-214 | 1 | 380 | 1 | 380 | |
| 4.85e-213 | 1 | 380 | 1 | 382 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6.44e-199 | 21 | 380 | 1 | 359 | Pectin Lyase A [Aspergillus niger] |
|
| 3.04e-197 | 21 | 380 | 1 | 359 | Pectin Lyase A [Aspergillus niger],1IDJ_B Pectin Lyase A [Aspergillus niger] |
|
| 1.59e-192 | 22 | 379 | 2 | 359 | Pectin Lyase B [Aspergillus niger] |
|
| 4.24e-07 | 149 | 312 | 107 | 273 | Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O88_A Chain A, Pectate Lyase C [Dickeya chrysanthemi],1O8D_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8E_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8F_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8G_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8H_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8I_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8J_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8K_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8L_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1O8M_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi],1PLU_A Chain A, Protein (pectate Lyase C) [Dickeya chrysanthemi],2PEC_A Chain A, PECTATE LYASE C [Dickeya chrysanthemi] |
|
| 1.00e-06 | 149 | 312 | 107 | 273 | Chain A, Pectate lyase C [Dickeya chrysanthemi] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.84e-262 | 1 | 380 | 1 | 380 | Probable pectin lyase A OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=pelA PE=3 SV=1 |
|
| 1.26e-216 | 1 | 380 | 1 | 378 | Probable pectin lyase A OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=pelA PE=3 SV=2 |
|
| 2.53e-216 | 1 | 380 | 1 | 378 | Probable pectin lyase A OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=pelA PE=3 SV=1 |
|
| 2.53e-216 | 1 | 380 | 1 | 378 | Probable pectin lyase B OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=pelB PE=3 SV=1 |
|
| 1.07e-215 | 1 | 377 | 1 | 377 | Pectin lyase A OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=pelA PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
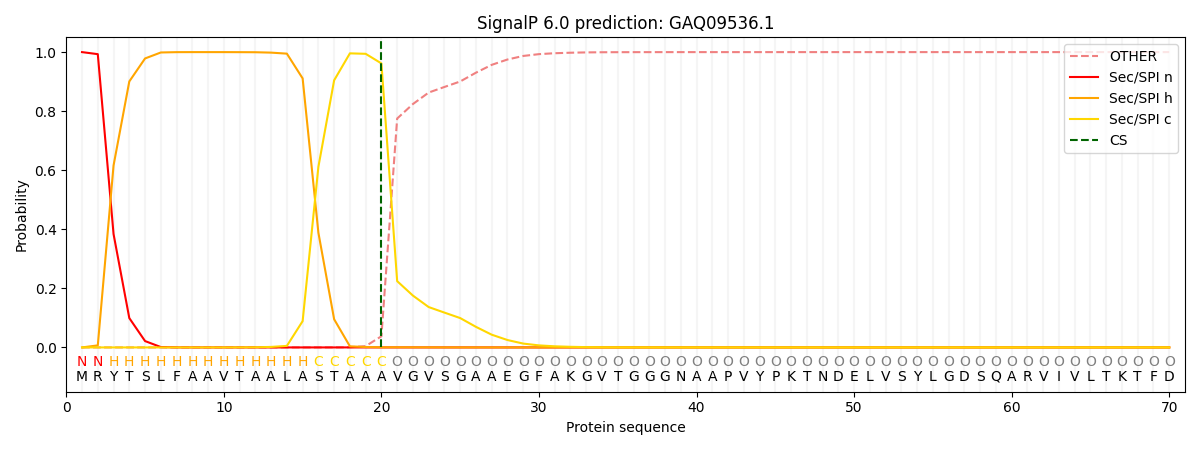
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000258 | 0.999713 | CS pos: 20-21. Pr: 0.9617 |