You are browsing environment: FUNGIDB
CAZyme Information: GAQ06773.1
You are here: Home > Sequence: GAQ06773.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus lentulus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus lentulus | |||||||||||
| CAZyme ID | GAQ06773.1 | |||||||||||
| CAZy Family | GH2 | |||||||||||
| CAZyme Description | probable acetylxylan esterase A | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.1.1.72:7 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE1 | 43 | 277 | 6.6e-23 | 0.9251101321585903 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 273828 | esterase_phb | 4.85e-40 | 46 | 249 | 1 | 199 | esterase, PHB depolymerase family. This model describes a subfamily among lipases of the ab-hydrolase family. This subfamily includes bacterial depolymerases for poly(3-hydroxybutyrate) (PHB) and related polyhydroxyalkanoates (PHA), as well as acetyl xylan esterases, feruloyl esterases, and others from fungi. [Fatty acid and phospholipid metabolism, Degradation] |
| 226040 | LpqC | 4.36e-26 | 13 | 289 | 18 | 277 | Poly(3-hydroxybutyrate) depolymerase [Secondary metabolites biosynthesis, transport and catabolism]. |
| 402228 | Esterase_phd | 2.14e-21 | 48 | 236 | 5 | 187 | Esterase PHB depolymerase. This family of proteins include acetyl xylan esterases (AXE), feruloyl esterases (FAE), and poly(3-hydroxybutyrate) (PHB) depolymerases. |
| 395257 | Peptidase_S9 | 6.55e-08 | 83 | 173 | 10 | 99 | Prolyl oligopeptidase family. |
| 224423 | DAP2 | 1.24e-07 | 46 | 172 | 382 | 507 | Dipeptidyl aminopeptidase/acylaminoacyl peptidase [Amino acid transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 3.02e-162 | 1 | 302 | 1 | 300 | |
| 2.24e-161 | 7 | 303 | 8 | 307 | |
| 2.24e-161 | 7 | 303 | 8 | 307 | |
| 2.24e-161 | 7 | 303 | 8 | 307 | |
| 2.24e-161 | 7 | 303 | 8 | 307 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.42e-155 | 28 | 302 | 1 | 275 | Acetyl xylan esterase from Aspergillus awamori [Aspergillus awamori],5X6S_B Acetyl xylan esterase from Aspergillus awamori [Aspergillus awamori] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.18e-173 | 1 | 302 | 1 | 307 | Probable acetylxylan esterase A OS=Aspergillus clavatus (strain ATCC 1007 / CBS 513.65 / DSM 816 / NCTC 3887 / NRRL 1 / QM 1276 / 107) OX=344612 GN=axeA PE=3 SV=1 |
|
| 1.25e-166 | 1 | 302 | 1 | 311 | Probable acetylxylan esterase A OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=axeA PE=3 SV=1 |
|
| 3.98e-162 | 7 | 303 | 8 | 307 | Probable acetylxylan esterase A OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=axeA PE=3 SV=1 |
|
| 1.14e-161 | 7 | 303 | 8 | 307 | Acetylxylan esterase A OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=axeA PE=1 SV=1 |
|
| 1.14e-161 | 7 | 303 | 8 | 307 | Probable acetylxylan esterase A OS=Aspergillus flavus OX=5059 GN=axeA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
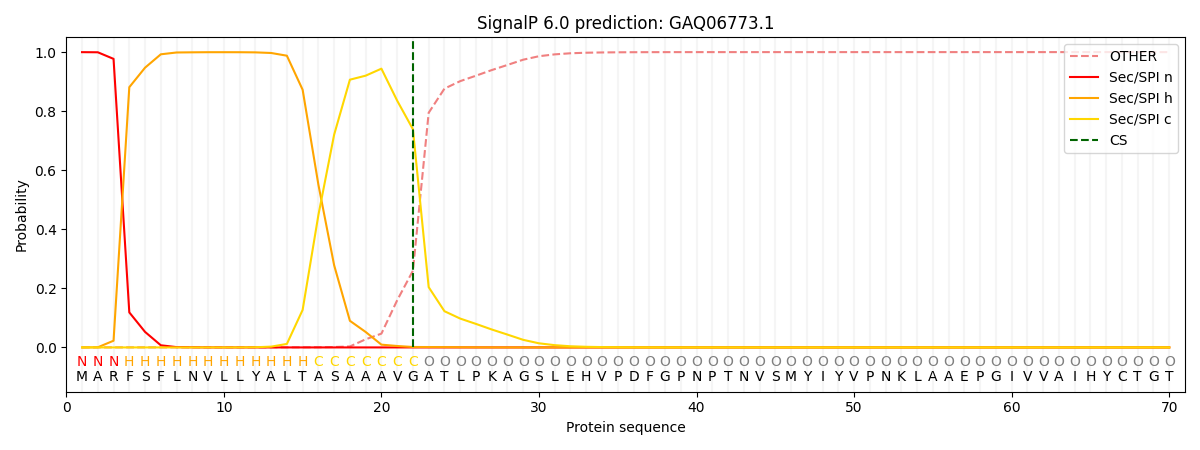
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.001071 | 0.998915 | CS pos: 22-23. Pr: 0.7388 |
