You are browsing environment: FUNGIDB
CAZyme Information: FVEG_13550-t26_1-p1
You are here: Home > Sequence: FVEG_13550-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Fusarium verticillioides | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Nectriaceae; Fusarium; Fusarium verticillioides | |||||||||||
| CAZyme ID | FVEG_13550-t26_1-p1 | |||||||||||
| CAZy Family | GT20 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 2753799; End:2757209 Strand: + | |||||||||||
Full Sequence Download help
| MIDMMDSGTS RRSVSMSGAV INGSIRHVTG LSLGASTPVP VVAIIGDKGS GKTSLFEALS | 60 |
| GLSLHGSRLS STKFPIHAYF THSNRTGTTL RARIRPGKVD GQNEELSDHL QSFEVNYEGD | 120 |
| LAAVNLPQIV NQATNHMQIA TAFSDEKEQE EQQVSDNVLV IEISGPDFFN MEIVDTPGLS | 180 |
| SSPAVDDTQE VTRSLIMDLV RQPNTLILSV VDATSDMSTQ EGPRLARAAD VTGNRIVAAV | 240 |
| TKCDLVPPEE RRFVLRAMRN KERHLGHPWF AIRSLSSNER DQDCTLETRN SVEDAFFSEM | 300 |
| EWSVLKRRRR VGAPSLKDYL KTRVNVLLRG SIELLTTILN TPSHSVAPSH APSITEKVQQ | 360 |
| TNIYAVSQET VVADVDLEFA SDAPRPTPLL NAVGVGLTVA ICLCLIGLGC SSMAVEIASE | 420 |
| GHWYRLFLLV TAPPQVILSL FFCQAIVVTI FQMIGPVSQL KVNSKFYSAK RTKRLDGTKV | 480 |
| LLPHVTIQCP VYKEGLEGVI RPTVQSLSAA IRNYESFGGT ASIFINDDGM QLVSEEEAEE | 540 |
| RRMFYKENNI GWVARPGHNV NGEGIKRFIR RGKFKKASNM NYALGISLKV EDKLVQIDRH | 600 |
| EKWTQVDEDA AYQKCLDEVL SEELGRAWVG GNIRVGDYLL LVDSDTRVPK DCLIDAVSEL | 660 |
| EASPQVAILQ FTSGVMNVTT SYFESGITFF TNLIYTAITY AVANGDVSPF VGHNAFIRWS | 720 |
| ALQEVGSQDG GIINGEQDPN ATTPYEKFWS ESHVSEDFDI SLRLQTLGYH IRLGAYCGDG | 780 |
| FKEGVSLTVY DELARWEKYA YGCSELLFWP IKDWWRHGIF TPLFKRFVMS NMPLGSKITI | 840 |
| MSYIGTYYAI GYSWIGSLLN YFLIGWLNGE LDHYYMSSWR VWVALVVVFS IAGNITLALI | 900 |
| RYRSQQVSLL KELWTCFKWV ALMFIFLGGI SMHVCKAIMC HMLSIDISWG ATSKEVEDTN | 960 |
| FFQEISIIIG GFKYVFIFCL AVTALMICGV YAFPYLWRID ELVAIFPLGT VIFCHFFLPI | 1020 |
| ALNPNLMKFT W | 1031 |
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT2 | 638 | 863 | 1.8e-47 | 0.9695431472081218 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 206738 | DLP_1 | 4.36e-47 | 38 | 320 | 2 | 265 | Dynamin_like protein family includes dynamins and Mx proteins. The dynamin family of large mechanochemical GTPases includes the classical dynamins and dynamin-like proteins (DLPs) that are found throughout the Eukarya. These proteins catalyze membrane fission during clathrin-mediated endocytosis. Dynamin consists of five domains; an N-terminal G domain that binds and hydrolyzes GTP, a middle domain (MD) involved in self-assembly and oligomerization, a pleckstrin homology (PH) domain responsible for interactions with the plasma membrane, GED, which is also involved in self-assembly, and a proline arginine rich domain (PRD) that interacts with SH3 domains on accessory proteins. To date, three vertebrate dynamin genes have been identified; dynamin 1, which is brain specific, mediates uptake of synaptic vesicles in presynaptic terminals; dynamin-2 is expressed ubiquitously and similarly participates in membrane fission; mutations in the MD, PH and GED domains of dynamin 2 have been linked to human diseases such as Charcot-Marie-Tooth peripheral neuropathy and rare forms of centronuclear myopathy. Dynamin 3 participates in megakaryocyte progenitor amplification, and is also involved in cytoplasmic enlargement and the formation of the demarcation membrane system. This family also includes interferon-induced Mx proteins that inhibit a wide range of viruses by blocking an early stage of the replication cycle. Dynamin oligomerizes into helical structures around the neck of budding vesicles in a GTP hydrolysis-dependent manner. |
| 404513 | Glyco_trans_2_3 | 2.51e-43 | 638 | 855 | 1 | 193 | Glycosyl transferase family group 2. Members of this family of prokaryotic proteins include putative glucosyltransferases, which are involved in bacterial capsule biosynthesis. |
| 395279 | Dynamin_N | 4.26e-17 | 42 | 242 | 1 | 167 | Dynamin family. |
| 224136 | BcsA | 3.77e-16 | 426 | 957 | 1 | 427 | Glycosyltransferase, catalytic subunit of cellulose synthase and poly-beta-1,6-N-acetylglucosamine synthase [Cell motility]. |
| 197491 | DYNc | 4.11e-12 | 32 | 246 | 17 | 205 | Dynamin, GTPase. Large GTPases that mediate vesicle trafficking. Dynamin participates in the endocytic uptake of receptors, associated ligands, and plasma membrane following an exocytic event. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CCT73007.1|GT0 | 0.0 | 16 | 1031 | 1 | 1016 |
| QGI99503.1|GT0 | 0.0 | 16 | 1031 | 1 | 1034 |
| QGI68613.1|GT0 | 0.0 | 16 | 1031 | 1 | 1034 |
| QKD62182.1|GT0 | 0.0 | 137 | 1031 | 1 | 912 |
| QGI85813.1|GT0 | 0.0 | 370 | 1031 | 5 | 642 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4P4S_A | 1.21e-13 | 39 | 335 | 13 | 285 | GMPPCP-bound stalkless-MxA [Homo sapiens] |
| 4P4T_A | 1.63e-13 | 39 | 363 | 34 | 326 | GDP-bound stalkless-MxA [Homo sapiens] |
| 4P4U_A | 1.68e-13 | 39 | 363 | 34 | 326 | Nucleotide-free stalkless-MxA [Homo sapiens] |
| 4P4S_B | 1.93e-13 | 39 | 335 | 28 | 300 | GMPPCP-bound stalkless-MxA [Homo sapiens] |
| 5UOT_0 | 1.05e-12 | 39 | 335 | 26 | 297 | CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_1 CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_2 CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_3 CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_4 CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_5 CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_6 CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_7 CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_8 CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_9 CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_A CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_B CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_C CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_D CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_E CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_F CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_G CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_H CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_I CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_J CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_K CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_L CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_M CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_N CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_O CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_P CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_Q CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_R CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_S CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_T CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_U CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_V CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_W CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_X CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_Y CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_Z CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_a CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_b CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_c CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_d CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_e CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_f CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_g CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_h CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_i CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_j CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_k CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_l CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_m CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_n CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_o CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_p CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_q CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_r CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_s CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_t CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_u CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_v CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_w CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_x CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_y CryoEM structure of the helical assembly of full length MxB [Homo sapiens],5UOT_z CryoEM structure of the helical assembly of full length MxB [Homo sapiens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| sp|P18588|MX1_RAT | 5.80e-15 | 39 | 347 | 61 | 344 | Interferon-induced GTP-binding protein Mx1 OS=Rattus norvegicus OX=10116 GN=Mx1 PE=1 SV=1 |
| sp|P09922|MX1B_MOUSE | 2.22e-14 | 39 | 347 | 36 | 320 | Interferon-induced GTP-binding protein Mx1 OS=Mus musculus OX=10090 GN=Mx1 PE=1 SV=1 |
| sp|A7VK00|MX2_PIG | 2.51e-14 | 39 | 335 | 115 | 385 | Interferon-induced GTP-binding protein Mx2 OS=Sus scrofa OX=9823 GN=MX2 PE=2 SV=1 |
| sp|Q6PW23|MX_EPICO | 1.15e-13 | 39 | 320 | 33 | 290 | Interferon-induced GTP-binding protein Mx OS=Epinephelus coioides OX=94232 GN=mx PE=2 SV=1 |
| sp|Q4U4N4|MX2_ICTPU | 1.51e-13 | 39 | 320 | 32 | 289 | Interferon-induced GTP-binding protein Mx2 OS=Ictalurus punctatus OX=7998 GN=mx2 PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
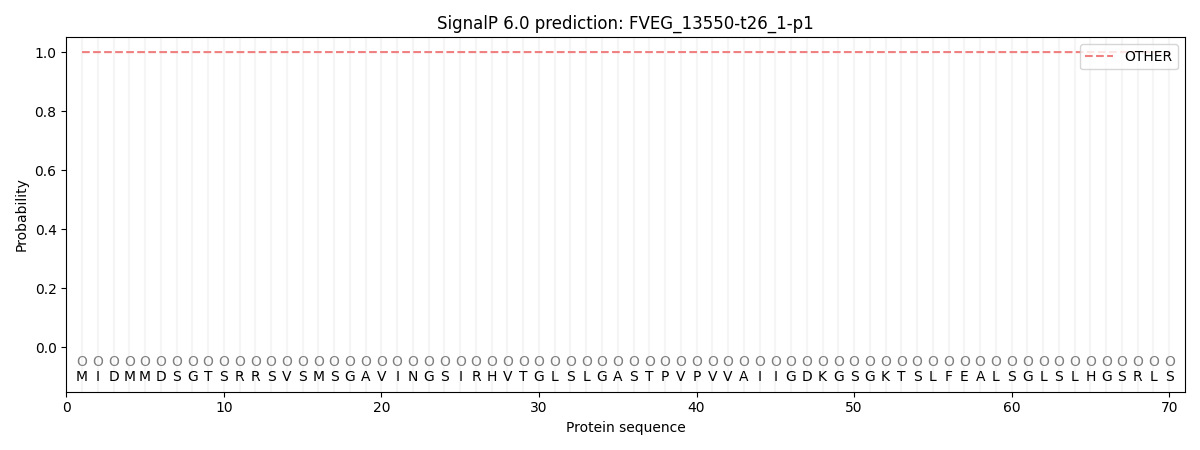
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000046 | 0.000000 |
TMHMM Annotations download full data without filtering help
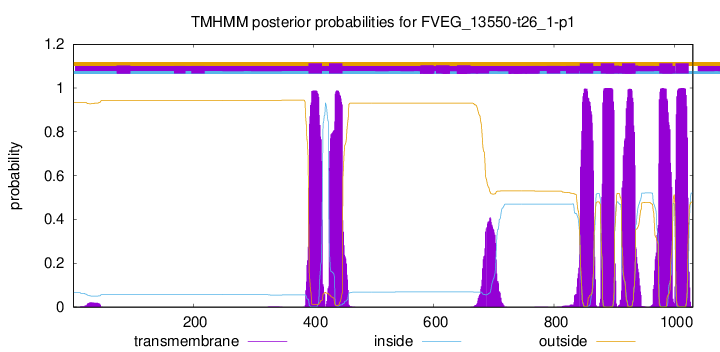
| Start | End |
|---|---|
| 392 | 414 |
| 426 | 448 |
| 843 | 865 |
| 878 | 900 |
| 913 | 935 |
| 975 | 997 |
| 1002 | 1024 |
